AbstractImproved methods for detection of Cryptosporidium oocysts in environmental and clinical samples are urgently needed to improve detection of cryptosporidiosis. We compared the sensitivity of 7 PCR primer sets for detection of Cryptosporidium parvum. Each target gene was amplified by PCR or nested PCR with serially diluted DNA extracted from purified C. parvum oocysts. The target genes included Cryptosporidium oocyst wall protein (COWP), small subunit ribosomal RNA (SSU rRNA), and random amplified polymorphic DNA. The detection limit of the PCR method ranged from 103 to 104 oocysts, and the nested PCR method was able to detect 100 to 102 oocysts. A second-round amplification of target genes showed that the nested primer set specific for the COWP gene proved to be the most sensitive one compared to the other primer sets tested in this study and would therefore be useful for the detection of C. parvum.
Cryptosporidium parvum (Apicomplexa: Cryptosporididae) is an intracellular parasitic protozoan that has emerged as an important cause of diarrhea among humans and animals [1]. In particular, the infection is more serious in immunocompromised patients and can become chronic and sometimes fatal [2]. Diagnosis is generally based on microscopic detection of oocysts, but this offers no information on the infected species and presents a challenge even to the most highly trained laboratory technician [3]. Therefore, a rapid, specific, and sensitive method is necessary for the detection of C. parvum. Molecular detection techniques, such as PCR-based methods, offer many advantages over microscopic methods. Several target genes have been previously reported to be useful for PCR detection of C. parvum [4-9]. In the present study, the sensitivity of various PCR target gene primer sets for C. parvum detection was compared.
Oocysts of C. parvum (KKU isolate) were maintained in specific pathogen-free C57BL female mice after immunosuppression by dexamethasone phosphate disodium salt (Sigma, St. Louis, Missouri, USA) provided ad libitum in drinking water at a dosage of 10 mg/ml [10]. Oocyst purification was carried out according to a method described by Petry et al. [11]. Purified oocysts were surface-sterilized by being placed in 10% sodium hypochlorite solution for 10 min and kept for < 2 week at 4℃ in filtered (0.22 µm) distilled water for the experiment.
For extraction of C. parvum DNA, 107 isolated oocysts were resuspended in the ASL lysis buffer included in the QIAquick stool mini kit (QIAGEN Inc, Valencia, California, USA). The samples were incubated at 70℃ for 30 min. The procedure was executed in accordance with the manufacture's recommendations. The extracted DNA was used as a template for PCR. All primer sets specific to the C. parvum target genes used in this study have been described previously by other authors (Table 1). The location of primer sequence in each specific C. parvum gene was shown in Fig. 1. The primer pair, cowpnest-F1 and cowpnest-R2, were designed for nested PCR specific for the COWP gene. PCR was performed with 4 primer sets, including Cry-15, Cry-19 (COWP), BcowpF, BcowpR (COWP-1), rRNAF, rRNAR (SSU rRNA), SB012F, and SB012R (random amplified polymorphic DNA). PCR amplification was performed in a 50 µl volume containing the template DNA (5 µl of genomic C. parvum DNA at a concentration equivalent to the number of C. parvum oocysts diluted from 105 oocysts), 10 mM Tris-HCl (pH 8.3), 2.5 mM MgCl2, 200 µM each of dATP, dCTP, dGTP, and dTTP, 25 pmole of each primer, and 2.5 U Taq DNA polymerase (Promega, Madison, Wisconsin, USA), respectively. All amplifications were performed in a Perkin-Elmer DNA thermal cycler (Model: 2400, Wellesley, Massachusetts, USA) with an initial denaturation at 94℃ for 5 min followed by 30 cycles of denaturation for 50 sec at 94℃, and annealing for 30 sec at the designated temperature (Table 1), extension for 50 sec at 72℃, and with a final extension at 72℃ for 10 min. For nested PCR, 2 µl of purified initial PCR product was used as a template. The nested PCR cycling conditions were identical to those used for PCR amplification, except that the annealing of the nested primers was performed at each temperature, as mentioned in Table 1. Amplified DNA fragments were analyzed by electrophoresis in a 1.5% (w/v) agarose gel stained with ethidium bromide (0.5 µg/ml) and visualized under a UV light system (Vilber Lourmat, France).
From the PCR amplification, we obtained the following products of the predicted size: 550 bp (COWP gene), 769 bp (COWP gene-1), 1,325 bp (SSU rRNA gene), and 433 bp (random amplified polymorphic DNA) (Fig. 2). COWP gene-1 was the larger PCR product than the COWP fragment amplified by the primers Cry-15 and Cry-19, and it included these regions. We also acquired the following predicted PCR products by nested PCR amplification: 311 bp (COWP gene), 550 bp (COWP gene-1), and 826 bp (SSU rRNA gene) (Fig. 2).
The lower detection limit of PCR amplification was 103-104 oocysts for all tested primer sets. However, a greater level of sensitivity was observed from the nested PCR amplification, in which the detection limit was as low as a single oocyst (Fig. 2; Table 2). The most sensitive nested PCR target gene was COWP, and the primers cowpnest-F1 and cowpnest-R2 could detect a single oocyst. The detection limit for the random amplified polymorphic DNA was 10,000 oocysts using the primers, SB012F and SB012R, for PCR amplification (Fig. 2D). Nested PCR was not performed with this primer set.
Although the stool microscopy was useful for identification of infection, this tool is relatively insensitive when small numbers of oocysts are excreted or the period of oocyst shedding is short. Therefore, many infections may escape microscopic detection [12]. The PCR method is becoming more widely used to detect pathogenic microorganisms, although this assay often requires multiple steps to achieve suitable DNA template preparations. So far, many PCR primers using various target genes have been constructed to detect Cryptosporidium, but the sensitivity and specificity were highly variable. The detection limits reported for PCR based methods by different authors have ranged from 1 to 10 oocysts in fecal samples of humans and calves, and in water sample [3-5,10], 5 to 50 oocysts in diluted bovine feces and in seeded concentrated environmental water samples [13-14], and 100 to 1,000 oocysts per gram of human feces or water [15-18].
In our study, nested PCR reactions specific for the COWP gene had an apparent advantage in sensitivity over other primers. Particularly, the combination of the primers, cowpnest F1 and cowpnest R2, was highly sensitive and could detect as little as 1 oocyst. Another primer pair for the COWP gene, BcowpF and BcowpR, was designed to produce a larger fragment of this gene to include the region amplified by cowpnest-F1 and cowpnest-R2. The sensitivity of nested PCR using this primer set was 100-fold lower than for the smaller product of the COWP gene, obtained using cowpnest-F1 and cowpnest-R2.
In our study, the number of oocysts used to produce the template DNA was calculated indirectly using diluted DNA extracted from purified oocysts. Therefore, the sensitivity limit may be different when this PCR method is applied to environmental or clinical samples. It has been demonstrated that substances present in feces, such as hemoglobin degradation products, bilirubin, and bile acids, may inhibit DNA amplification and lead to false-negative PCR results, as described by Lantz et al. [19]. In addition, inefficient methods for isolating small numbers of the organism and resistance of the organism to disruption and lysis could lower the sensitivity [4].
In a previous study, Wu et al. [5] demonstrated high sensitivity of the PCR approach, detecting a single C. parvum oocyst using the primers SB012F and SB012R. However, in the present study, the detection sensitivity of this primer set was 1,000-fold lower than in the previous report. The reason for this discrepancy is unclear.
In conclusion, we developed a highly sensitive primer set specific to the COWP gene that can be used for detection of C. parvum. This highly sensitive nested PCR method will be helpful for facilitating the detection of cryptosporidiosis in patients with a low number of oocysts and could be a valuable tool for detection of oocysts in environmental and clinical samples.
REFERENCES1. Clark DP. New insights into human cryptosporidiosis. Clin Microbiol Rev 1999;12:554-563. PMID: 10515902.
2. Tzipori S, Ward H. Cryptosporidiosis: biology, pathogenesis and disease. Microbes Infect 2002;4:1047-1058. PMID: 12191655.
3. Coupe S, Sarfati C, Hamane S, Derouin F. Detection of Cryptosporidium and identification to the species level by nested PCR and restriction fragment length polymorphism. J Clin Microbiol 2005;43:1017-1023. PMID: 15750054.
4. Johnson DW, Pieniazek NJ, Griffin DW, Misener L, Rose JB. Development of a PCR protocol for sensitive detection of Cryptosporidium oocysts in water samples. Appl Environ Microbiol 1995;61:3849-3855. PMID: 8526496.
5. Wu Z, Nagano I, Matsuo A, Uga S, Kimata I, Iseki M, Takahashi Y. Specific PCR primers for Cryptosporidium parvum with extra high sensitivity. Mol Cell Probes 2000;14:33-39. PMID: 10722790.
6. Xiao L, Escalante A, Yang C, Sulaiman I, Escalante AA, Montali RJ, Fayer R, Lal AA. Phylogenetic analysis of Cryptosporidium parasites based on the small-subunit rRNA gene locus. Appl Environ Microbiol 1999;65:1578-1583. PMID: 10103253.
7. Spano F, Puri C, Ranucci L, Putignani L, Crisanti A. Cloning of the entire COWP gene of Cryptosporidium parvum and ultrastructural localization of the protein during sexual parasite development. Parasitology 1997;114:427-437. PMID: 9149414.
8. Pedraza-Diaz S, Amar C, McLauchlin J. The identification and characterization of an unusual genotype of Cryptosporidium from human faeces as Cryptosporidium meleagridis. FEMS Microbiol Lett 2000;189:189-194. PMID: 10930736.
9. Xiao L, Limor J, Morgan UM, Sulaiman IM, Thompson RC, Lal AA. Sequence differences in the diagnostic target region of the oocyst wall protein gene of Cryptosporidium parasites. Appl Environ Microbiol 2000;66:5499-5502. PMID: 11097936.
10. Yang S, Healey MC. The immunosuppressive effects of dexamethasone administered in drinking water to C57BL/6N mice infected with Cryptosporidium parvum. J Parasitol 1993;79:626-630. PMID: 8331488.
11. Petry F, Robinson HA, McDonald V. Murine infection model for maintenance and amplification of Cryptosporidium parvum oocysts. J Clin Microbiol 1995;33:1922-1924. PMID: 7665672.
12. McCluskey BJ, Greiner EC, Donovan GA. Patterns of Cryptosporidium oocyst shedding in calves and a comparison of two diagnostic methods. Vet Parasitol 1995;60:185-190. PMID: 8747901.
13. Webster KA, Smith HV, Giles M, Dawson L, Robertson LJ. Detection of Cryptosporidium parvum oocysts in faeces: comparison of conventional coproscopical methods and the polymerase chain reaction. Vet Parasitol 1996;61:5-13. PMID: 8750678.
14. Rochelle PA, De Leon R, Stewart MH, Wolfe RL. Comparison of primers and optimization of PCR conditions for detection of Cryptosporidium parvum and Giardia lamblia in water. Appl Environ Microbiol 1997;63:106-114. PMID: 8979344.
15. Gobet P, Buisson JC, Vagner O, Naciri M, Grappin M, Comparot S, Harly G, Aubert D, Varga I, Camerlynck P. Detection of Cryptosporidium parvum DNA in formed human feces by a sensitive PCR-based assay including uracil-N-glycosylase inactivation. J Clin Microbiol 1997;35:254-256. PMID: 8968918.
16. Mayer CL, Palmer CJ. Evaluation of PCR, nested PCR, and fluorescent antibodies for detection of Giardia and Cryptosporidium species in wastewater. Appl Environ Microbiol 1996;62:2081-2085. PMID: 8787406.
17. Balatbat AB, Jordan GW, Tang YJ, Silva JJ. Detection of Cryptosporidium parvum DNA in human feces by nested PCR. J Clin Microbiol 1996;34:1769-1772. PMID: 8784586.
18. Lee SU, Joung M, Ahn MH, Huh S, Song H, Park WY, Yu JR. CP2 gene as a useful viability marker for Cryptosporidium parvum. Parasitol Res 2008;102:381-387. PMID: 18060431.
Fig. 1Position of the primer pair specific to Cryptosporidium parvum gene for PCR and nested PCR. Target genes of C. parvum for PCR were found at GenBank accession no. (COWP, Z22537; SSU rRNA, AF093489; SB012, AF161076). 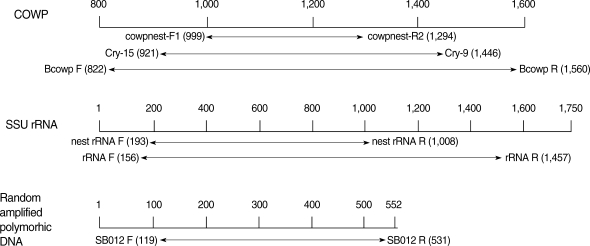
Fig. 2Amplification by PCR and nested PCR of Cryptosporidium parvum DNA extracted from purified C. parvum oocysts. The upper gel shows amplification by PCR, and the lower gel shows amplification by nested PCR using amplified PCR products as template DNA. M, 100 bp DNA ladder marker (Promega); (A) COWP gene; (B) COWP-1 gene; (C) SSU rRNA gene; (D) SB012 (random amplified polymorphic DNA). 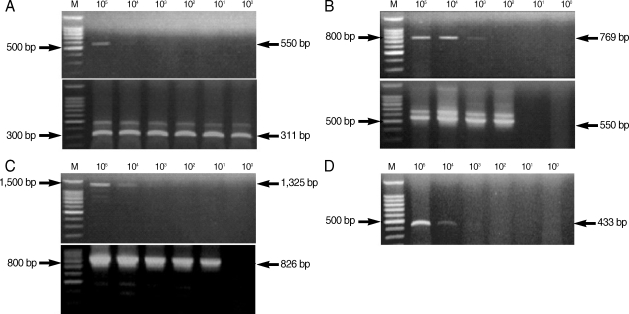
Table 1.Primer sets for PCR and nested PCR specific to C. parvum genes
Table 2.Differences of sensitivity among various primer sets for PCR or nested PCR of C. parvum
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||