INTRODUCTION
Acanthamoeba, which cause granulomatous amebic encephalitis (GAE) and amebic keratitis, comprise of 2 distinct stages, trophozoite and cyst. The cyst is formed from a trophozoite when desiccation, starvation, or other adverse conditions prevail, and conversely it transforms to the trophozoite when the conditions are favorable [1]. During encystation of the amoeba, various changes occur in cellular composition, and formation of autophagosome is one of the changes [2]. Autophagy, the molecular machinery for self-eating [3], is necessary for the recycling of cytoplasmic components for survival under a starvation condition [4]. Two ubiquitin-like conjugation systems, autophagy-related protein (Atg) 12 system and Atg8 system, are necessary for autophagosome formation [5]. Analogous to ubiquitination, Atg12 is conjugated to Atg5 by Atg7, an E1-like protein, and Atg10, an E2-like protein [6]. Similarly, Atg7 and Atg3 are the respective E1-like and E2-like proteins that mediate the conjugation of Atg8 localization to phosphatidylethanolamine (PE) [6]. Studies in Saccharomyces cerevisiae have identified 18 ATG genes required for autophagosome formation, and most are also found in higher eukaryotes [7].
Recently, we have studied the role of autophagy system in encysting Acanthamoeba mainly focusing on the Atg8-PE conjugation system. According to the results from other organisms, the ubiquitin-like protein Atg8 is cleaved by cysteine protease Atg4, producing Atg8Gly-116 [8], and transferred to Atg3 after being activated by Atg7 [9,10]. Finally, Atg3 conjugates Atg8 with PE [10] and then Atg8-PE is bound to the autophagosome membrane.
Picazarri et al. [2] reported that Entamoeba invadens possess the Atg8 conjugation system, consisting of Atg8, Atg4, Atg3, and Atg7, but lack the Atg5-to-Atg12 conjugation system. In Trypanosoma brucei, the crystal structure of TbAtg8 was reported [11], and autophagy pathway of Leishmania major was monitored via the expression of the autophagosome marker GFP-Atg8 [12]. Otto et al. [13] observed that apg5 and apg7 are essential for the development of Dictyostelium discoideum.
In Acanthamoeba, only Atg8 within the Atg8 conjugation system was identified [14], and the other members, including Atg4, Atg7, and Atg3, and the relationship among the members have yet to be elucidated. In this study, we identified and characterized autophagy-related protein 3 from Acanthamoeba castellanii (AcAtg3). Furthermore, we verified that Atg8 lipidation is mediated by Atg3 in encysting Acanthamoeba.
MATERIALS AND METHODS
Amoeba cultivation
A. castellanii Castellani trophozoites were obtained from the American Type Culture Collection (ATCC 30011) and were grown axenically in PYG (peptone-yeast-glucose) medium (10 g proteose peptone, 10 g yeast extract, 10 ml of 50% glucose, 10 ml of 0.5 M Na2HPO4, and 10 ml of 0.5 M K2HPO4 in 970 ml distilled water with the final pH adjusted to 6.5) at 25℃ in an incubator (Sanyo, San Diego, California, USA). Encystment was induced by the procedure of Bowers and Korn [15]. Approximately 5×105 cells were washed once with an encystment medium, containing 95 mM NaCl, 5 mM KCl, 8 mM MgSO4, 0.4 mM CaCl2, 1 mM NaHCO3, 20 mM Tris-HCl, pH 9.0, and incubated in 10 ml of the encystment medium in cell culture dishes at 25℃ for 3 days.
Real-time PCR
Total RNA was purified using TRIzol Reagent (Gibco BRL, Rockville, Maryland, USA), and cDNA was synthesized using RevertAid™ First Strand cDNA synthesis kit (Fermentas, Hanover, Indiana, USA). Real-time PCR was performed using the GenAmp 5700 SDS (Biosystems, Barcelona, Spain) and the default thermocycler program for all genes: 10 min of pre-incubation at 95℃ followed by 40 cycles of 15 sec at 95℃ and 1 min at 60℃. Individual reactions were carried out in 20 µl volumes in a 96-well plate containing 1×buffer, 3.5 mM MgCl2, 0.2 mM dNTPs, different concentrations of sense and antisense primers (sense 5'-GCGCACGTACGATATCTCCATC and antisense 5'-ATGAACACTTGGTTCGGCGTC) 0.025 U/µl enzyme, and 1:66,000 SYBR Green (Bionics, Seoul, Korea). All reactions were made using SYBR Premix Ex Taq™ (Takara, Shiga, Japan). The reference gene was 18s rDNA (sense 5'-TCCAAT TTTCTGCCACCGAA and antisense 5'-ATCATTACCCTAGTCCTCGCGC).
Transient transfection of Acanthamoeba with Atg3
To investigate intracellular localization of AcAtg3, the gene was cloned into the pUb vector [16]. The AcAtg3 cDNA was PCR amplified with primers (sense 5'-GCGGAATTCATGACTTGGTGGGATAAGACA and antisense 5'-CGCACTAGTGCTCTCCATCTCCATGGTGTA) that included sites for Eco RI at the 5' end and Spe I at the 3' end, and were inserted into the pUb vector upstream of the gene for EGFP. This plasmid was transfected into viable A. castellanii. Transient transfection was performed by Superfect transfection reagent (Qiagen, Valencia, California, USA) as previously described [16].
Atg3 gene silencing by siRNA
Transfection of siRNA was performed for the confirmation of gene silencing of AcAtg3. Fluorescence-tagged siRNA was transfected to live Acanthamoeba, and transfection efficiency was observed under fluorescent microscopy. Small interfering RNA (siRNA) targeting the Atg3 gene of A. castellanii was synthesized by Sigma-Proligo (Boulder, Colorado, USA), based on the cDNA sequence of the gene. The siRNA duplex with sense (5'-[Flc]AGGCAAAUAAGACGCUCAU[dT][dT]) and anti-sense (5'-AUGAGCGUCUUAUUUGCCU[dT][dT]) sequences were used. The siRNA (4 µg) was transfected to A. castellanii trophozoites at a density of 4×105 cells. Transfection was performed by Superfect transfection reagent (Qiagen) as previously described [16]. As a negative control, siRNA with a scrambled sequence (Ambion, Austin, Texas, USA) absent in Acanthmaoeba was used.
Microscopy
Amoebae expressing EGFP were selected and allowed to adhere to a Falcon cell culture dish (BD, Piscataway, New Jersey, USA). The cells were observed using a LSM 5 EXCITER Scalable confocal system (Zeiss, Hamburg, Germany). EGFP- and LysoTracker Red DND 99-mediated fluorescence was achieved using band-pass filters that provided excitation and emission wavelengths of 500-530 nm and 570-590 nm, respectively.
Western blot analysis
For production of the antibody against A. castellanii Atg8 (AcAtg8) [14], the GST Gene Fusion System was utilized (Amersham Biosciences, Buckinghamshire, England). The PCR product of Atg8 was cloned into pGEX-4T-2 vector. GST-AcAtg8 fusion protein (>100 µg) of A. castellanii was subcutaneously injected into 6-week-old SD rats (Bionics, Seoul, Korea). Blood samples were collected from the rats before administering boosters, as to monitor antibody production. The specific antibody was confirmed after doing western blot of cell lysate. SDS-PAGE and immunoblotting were performed as described [17].
RESULTS
Cloning and characterization of AcAtg3
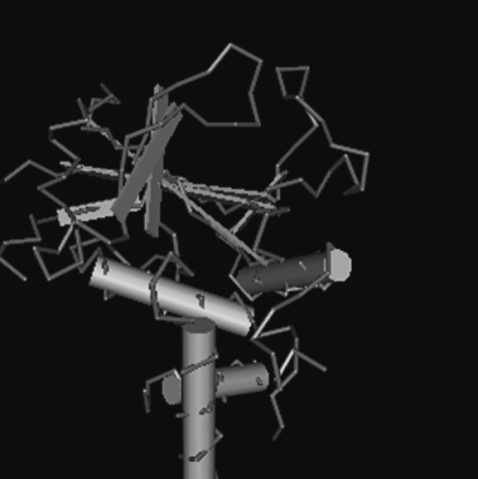
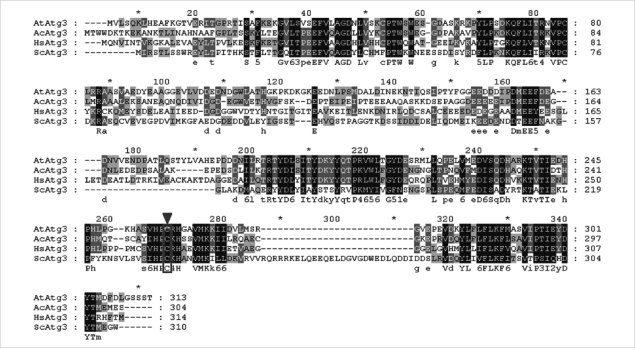
We identified the full-length open reading frame (ORF) of Atg3 in the encysting A. castellanii (AcAtg3) cDNA library (GenBank No. GU270859). This gene has a single ORF of 915 bp, predicting a protein of 304 amino acid residues. Based on homology searches, the deduced amino acid sequences of the AcAtg3 showed a conserved cysteine active site (Fig. 1). The predicted 3D structure of AcAtg3 was shown as a hammer-like shape determined by a specialized BLAST in NCBI that search for conserved domains with a protein sequence (Fig. 2). The topology of the head moiety of AcAtg3 is very similar to canonical E2 enzymes.
mRNA expression pattern of AcAtg3 during encystation
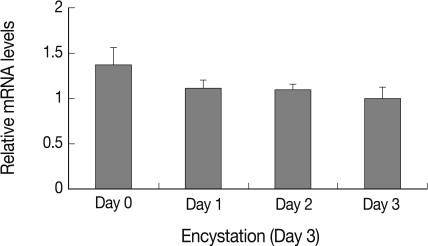
Expression pattern of AcAtg3 was investigated using real-time PCR on mRNA of trophozoite and cyst forms each day after induction of encystation. As shown in Fig. 3, level of AcAtg3 mRNA was similar or a little less than those of the trophozoites during encystation. It is supposed that AcAtg3 can be recycled during encystation in Acanthamoeba.
Intracellular localization of AcAtg3
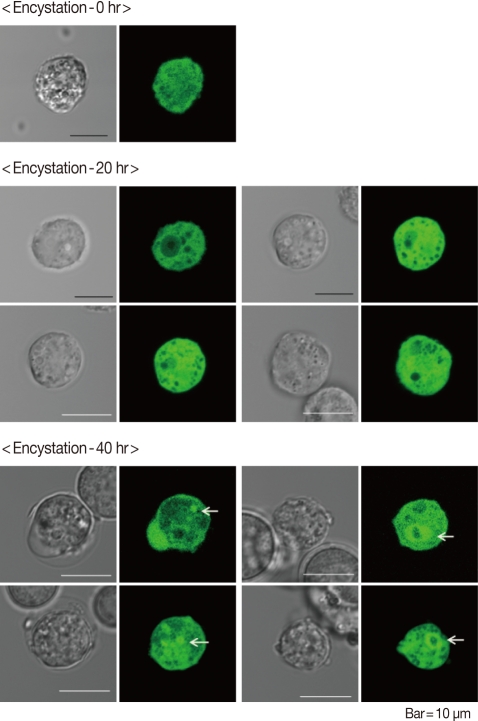
AcAtg3-EGFP fusion protein was transfected to live Acanthamoeba. The amoeba expressing Atg3 as an EGFP fusion protein showed dispersed fluorescence in its cytoplasm during the trophozoite and early stages of the encysting amoeba (Fig. 4; 0 hr and 20 hr). Interestingly, the localization was changed by encystation. After 40 hr of encystation induction, dispersed Atg3-EGFP fusion proteins were gathered and localized around ball-like structures (Fig. 4; 40 hr). We supposed that this structure could be an autophagosome. To confirm the localization of AcAtg3 as an autophagosome, transfected amoebae were stained with Lyso Tracker Red DND-99, an autophagosome marker. The position of the AcAtg3 co-localized or localized around the autophagosome membrane (Fig. 5). This result suggests that AcAtg3 is associated with the autophagosomal membranes in Acanthamoeba.
Effect of Atg3 siRNA on cyst formation and lipidation of Atg8 in Acanthamoeba
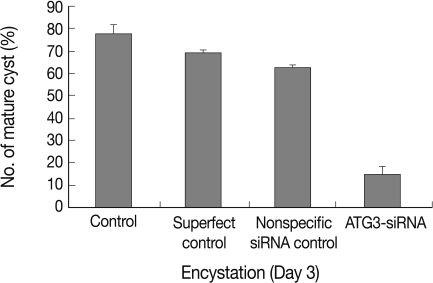
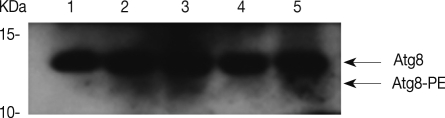
Transfection efficiency of siRNA was confirmed by FACS analysis. Of the treated trophozoites, 82% were detected as transfected cells (data not shown). Transfected cells were transferred to encystment media, and the number of mature cysts were compared with controls; superfect transfection reagent control, nonspecific siRNA control and AcAtg3-siRNA transfected cells (Fig. 6). The formation of mature cysts was almost completely inhibited in AcAtg3-siRNA transfected cells while the other control groups didn't show any differences. To search the reason why mature cysts were not formed in AcAtg3-siRNA transfected cells, western blot analysis using an AcAtg8 antibody was undertaken. The formation of Atg8-PE conjugate was investigated among trophozoite cells, encysting cells for 1 day and 2 days, and AcAtg3-siRNA transfected cells (encysting cells for 1 day and 2 days). Atg8-PE was present in encysting cells for 1 day and 2 days (lanes 2 and 3). However, it was absent in AcAtg3-siRNA transfected cells during encystation (lanes 4 and 5) (Fig. 7). This result shows that AcAtg3 plays a crucial role in AcAtg8-PE conjugation to form the autophagosome during encystation in Acanthamoeba.
DISCUSSION
In the present study, we have identified and characterized the role of Atg3 in A. castellanii (AcAtg3). The mRNA expression levels of AcAtg3 did not increase during encystation. However, transfected cells with AcAtg3-siRNA were not transformed into mature cysts (Fig. 6). AcAtg3-EGFP fusion protein was localized around the autophagosomal membranes (Fig. 5), and it was confirmed that AcAtg3 was associated with AcAtg8-PE conjugation during encystation in A. castellanii (Fig. 7).
Atg3 (also APG3 and PC3-96) is a ubiquitous member of the Atg3 family of proteins. The crystal structure of S. cerevisiae Atg3 is topologically similar to canonical E2 enzymes [18]. All E2 enzymes have a conserved E2 core composed of a central 4-stranded β-sheet and 4 α-helices surrounding it [19]. Atg3 functions as an E2-like enzyme during the initial stages of autophagosome formation by catalyzing the transfer of Atg7 bound Atg8 (known as LC3, GATE16, and GABA-RAP in mammals) to PE, the critical step for autophagy [20]. In Fig. 7, AcAtg8-PE conjugates were detected in lanes 2 and 3 (during encystation) with anti-Atg8. However, after the transfection of the AcAtg3-siRNA, AcAtg8-PE conjugates were not detected during encystation (lanes 4 and 5). It was confirmed that Atg3 has an activity for Atg8 lipidation for autophagosome formation in encysting Acanthamoeba. The expression pattern of Atg3/Apg3 has not yet been reported from other organisms under differentiation. In the tick Haemaphysalis longicornis, real-time PCR results revealed that the expression of HIAtg3, HIAtg4, and HIAtg8 showed higher levels during the non-feeding period rather than the feeding period. Their expression levels were higher in the midgut, a digestive organ, of unfed rather than fed adults [21]. These results suggested that the Atg8 conjugation system is at work in unfed ticks, and the starved condition appears to be associated with the increased expression of HIAtg genes in the midgut of unfed ticks [21]. We reported previously that the mRNA expression of AcAtg8 showed higher levels during encystation of Acanthamoeba [14]. Contrary to our previous results, the mRNA expression levels of AcAtg3 did not increase during encystation (Fig. 3). Under the condition of nutrient starvation in yeast cells, Atg3 protein is processed into a mature form within the vacuoles [18]. We therefore speculated that Atg3 of Acanthamoeba was appropriated in recycled forms during encystation. Fig. 5 shows the direct intracellular localization of AcAtg3 during encystation of Acanthamoeba. Evenly distributed Atg3-EGFP proteins throughout the cytoplasm in trophozoite cells were gathered around the autophagosomal membranes in encysting Acanthamoeba (Fig. 5, arrows). It is the first time that we determined the intracellular localization of Atg3 protein in encysting cells. It can be summarized that the AcAtg3 plays an important role in Atg8 lipidation, which takes place on the side of autophagosomal membranes.
Autophagy is a degradative pathway necessary for the clearance of damaged or superfluous proteins and organelles, and the recycling of intracellular constituents as well as providing energy during periods of metabolic stress or starvation [4]. Autophagy formation is a means of survival in encysting Acanthamoeba. If we understand the molecular mechanism of autophagy formation and disrupt the encysting process, we can protect the infection of cyst forming pathogenic protozoa. The results of this study could provide important information needed to understand the regulatory mechanism of autophagosome in cyst forming protozoa.