AbstractSince Kudoa septempuntata was identified as a causative agent of food poisoning associated with raw olive flounder Paralichthys olivaceus, interest and concern regarding the parasite have increased. However, there have been no investigations or reports of other Kudoa species infecting the fish (except for K. paralichthys, which infects the brain) in Korea. We found cysts filled with myxospores of Kudoa species in muscles of cultured olive flounder specimens and identified these to the species level. Mature spores were quadrate, measuring 8.7±0.5 μm in length, 9.2±0.4 μm in thickness, and 12.9±0.6 μm in width. The spores containing 4 polar capsules had a length of 2.1±0.2 μm and a width of 1.8±0.3 μm. The partial 18S and 28S rDNA of isolates showed 99–100% similarities with K. ogawai. Using these morphological and molecular analyses, the species was identified as K. ogawai. This study is the first report of K. ogawai infection in cultured olive flounder in Korea.
Myxozoans are parasites that complete a life cycle through vertebrate (mainly fish) and invertebrate (mainly annelid) hosts [1]. The genus Kudoa contains myxosporeans with 4 or more shall valves and polar capsules, and the parasites infect various fish species [2,3]. Some Kudoa spp. reduce the commercial value of fish by causing post-mortem liquefaction (such as K. thyrsites, K. lateolabracis, and K. neothunni) [3–7], cyst formation in muscles (K. iwatai and K. amamiensis) [8,9], and spinal deformation (K. yasunagai) [2,10,11]. In addition, previous studies have reported the relationship between Kudoa spp. and food-poisoning [12–14]. These findings have led to increased interest and concern regarding Kudoa infection in commercially important fish. Olive flounder, Paralichthys olivaceus, is an important fish species cultured in Korea, and there have been several reports of myxosporean infections (Enteromyxum leei, Parvicapsula anisocaudata, P. curvatura, and Sinuolinea capsularis), including Kudoa spp. (K. septempunctata and K. paralichthys), in the fish [15–19]. The Fish Vaccine Research Center has been monitoring the parasitic infections of olive flounder cultured in Jeju island and has isolated another Kudoa sp. from olive flounder muscle. The aim of the present study was to identify the Kudoa sp. using morphological and molecular analyses and to reveal differences between the present isolate and other Kudoa spp. known to infect olive flounder. In addition, the information obtained from this study will be applied to the inspection and identification of Kudoa spp.
Olive flounder samples (n=2, 56.4±4.3 cm) were obtained from an olive flounder farm situated on Jeju island. The fish samples were examined microscopically to detect K. septempunctata spores in squash preparations, and cysts filled with another Kudoa sp. were found. The cysts were collected using forceps and were squashed on a glass slide. The spores were wet mounted and observed under a light microscope and photographed at 400×or 1,000×magnification. Measurements of myxospores were obtain from 20 spores using the ImageJ image processing software (available at http://rsb.info.nih.gov/ij/) according to Lom and Arthur’s criteria (1989) [20].
The myxospores were collected and passed through a 40 μm cell strainer. DNA was extracted from partially purified parasites using the AccuPrep Genomic DNA Extraction Kit (Bioneer, Daejeon, Korea) following the manufacturer’s instructions. Portions of 18S and 28S rDNA were amplified by PCR using a combination of primers that we (28S-KO2000F: 5′-CTGTCCGTACCGAATCCG-3′) and other groups (18S-18e: 5′-CTGGTTGATCCTGCCAGT-3′, 18S-MyxospecF: 5′-TTCTGCCCTATCAACTWGTTG-3′, 18S-MyxospecR: 5′-GGTTTCN CDGRGGGMCCAAC-3′, 18S-ERIB10: 5′-CTTCCGCAGGTTCACCTA-3′, 28S-Kt28SF: 5′-CAAGACTACCTGCTGAAC-3′, 28S-NLF1050: 5′-AATCGAACCATCTAGTAGCTGG-3′, 28S-NLR1270: 5′-TTCATCCCGCATCGCCAGTTC-3′, 28S-NLR1694(mo): 5′-GTTAGGCAATGGCTTAGGACC-3′, 28S-3R: 5′-GAGCACTGGGCAGAAATC-3′, 28S-NLR3113(mo): 5′-GTCTAAACCCAGCTCACGTTC-3′, and 28S-NLR3284: 5′-TTCTGACTTAGAGGCGTTCAG-3′) have designed [2,16,21–25]. PCR products were treated with the AccuPrep Genomic PCR Purification Kit (Bioneer) to remove excess primers and dNTPs, and were directly sequenced using BigDyeTM Terminator v3.1 in an ABI 3730xl Sequencer. The 18S and 28S rDNA sequences of isolates were compared in the GenBank database using the Basic Local Alignment Search Tool (BLAST) search engine to find sequences with a high degree of similarity. Multiple alignments of 18S rDNA sequences were made using Clustal X 2.0 [26] with 7 homologous sequences from Kudoa spp. and one sequence from E. leei (outgroup) reported in olive flounder (except K. lateolabracis). The phylogenetic tree was constructed using the neighbor-joining method in MEGA 7 [27].
Oval cysts 1 mm in length approximately, were observed in the squashed muscle sample (Fig. 1A). Cysts were shorter in length and thicker than the pseudocysts of K. septempunctata that are usually detected, and the myxospores were also different in form from those of K. septempunctata (Fig. 1B, C). The myxospores were quadrate in the apical view and had 4 equal spore valves, each containing one polar capsule with a rounded peripheral edge. In addition, the appearance of the 4 polar capsules was similar to a 4-leaf clover (Fig. 1D). In side view, spores were rounded triangle-shaped, and prominent apical projections were observed (Fig. 1E). The spores measured 8.7±0.5 μm (8.3–9.6 μm) in length, 9.2±0.4 μm (8.4–10.0 μm) in thickness, and 12.9±0.6 μm (11.5–13.9 μm) in width. The polar capsule length was 2.1±0.2 μm (1.7–2.5 μm) and polar capsule width was 1.8±0.3 μm (1.4–2.5 μm) (Table 1). The species isolated in the present study was then compared with 8 Kudoa spp. that have been previously reported in olive flounder. Kudoa ogawai was the most similar to the isolated species in terms of measurements and shape of spores, number of valves per spore, apical projection, and infection site. Kudoa paralichthys and K. shiomitsui showed similarities to the isolate in terms of spore shape and number of valves per spore. However, they are smaller than the isolate and have a different infection site (brain and heart vs muscle). In addition, they have small projections that are not easily observed, while the isolate has distinct apical projections. Although K. lateolabracis and K. thyrsites had similarities number of valves per spore and share the same infection site as the isolate, they have distinct spore shapes with one large polar capsule. Kudoa septempunctata and K. igami have similar spore measurements and the same infection site as the isolate. However, they have a different spore shape (stellate vs. quadrate) as well as a different number of valves per spore (6 or 7 vs 4). In addition, their apical projections are not prominent as those of the isolate (Table 1) [4,11,19,28–30].
Partial sequences of the 18S rDNA (1,650 bp) and 28S rDNA (3,395 bp) were obtained from the isolate and had 100 and 99.9% similarities with the 18S (KX163082; 1,631/1,631) and 28S rDNA (KX163083; 3,318/3,323) sequences of K. ogawai isolated from olive flounder in Japan, respectively. Unfortunately, there is no 18S rDNA sequence of K. paralichthys available on GenBank. In addition, the 18S rDNA sequence of K. lateolabracis reported from olive flounder could not be found, thus the sequence of K. lateolabracis isolated from Chinese sea bass was used to construct the phylogenetic tree. The phylogenetic tree was divided into 4 groups and the isolate was clustered with K. ogawai (Fig. 2). Based on the morphological and molecular comparisons, the myxospores were identified as K. ogawai and the sequences were deposited in GenBank (accession numbers MK850343 and MK850346, respectively).
Korea and Japan have a food culture that involves eating raw fish, which increase the risk of food poisoning. Kudoa septempunctata has been reported as a causative agent of food poisoning after ingesting raw olive flounder [12,31]. Kawai et al. [12] reported that oral administration of K. septempunctata spores elicited accumulation of fluid in the intestine and produced watery diarrhea in suckling mice. They also showed that the direct feeding of olive flounder meat containing K. septempunctata spores as well as purified spores cause vomitting in house musk shrews. Ohnish et al. [31] revealed that K. septempunctata sporoplasm plays an important role in mediating the toxicity of K. septempunctata. In addition, previous studies have investigated the epidemiological relationship between K. septempunctata ingestion and food poisoning in Korea and Japan [13,32]. Iwashita et al. [32] reported 3 clinical cases related to K. septempunctata ingestion in the Mie prefecture of Japan, and Kim et al. [13] also reported on the epidemiologic characteristics of 16 outbreaks of foodborne illnesses caused by K. septempunctata in the Gyeonggi Province of Korea. However, some studies have suggested that there is no relationship between food poisoning and ingesting olive flounder infected by K. septempunctata [33–35]. Jang et al. [33] showed that suckling mice inoculated with 5×106–7 K. septempunctata spores did not show symptoms of diarrhea, and no pathological change in intestine of the mice. Ahn et al. [34] came to similar results in study using adult mice. In addition, they exhibited that K. septempunctata spores did not cause inflammation on Caco-2 cells [35]. It is therefore up for discussion whether K. septempunctata causes food poisoning or not, however a few cultured olive flounder are undeniably infected by the parasite.
In the present study, Kudoa sp. was discovered to infect the muscle of cultured olive flounder, and the isolate was identified as K. ogawai based on morphological and molecular comparisons. There have been no studies revealing the relationship between K. ogawai and food poisoning, however, there have been several reports that other myxosporeans (not K. septempunctata) may cause foodborne diseases [14,36,37]. Kawase et al. [36] reported that K. hexapunctata genes were detected in the feces of a patient exhibiting diarrhea and vomiting after eating raw juvenile Pacific bluefin tuna, Thunnus orientalis, and Ohnishi et al. [14] also detected the genes and spores of K. iwatai from Rock bream (barred knifejaw), Oplegnathus fasciatus, and Blackhead seabream, Acanthopagrus schlegelii, associated with gastrointestinal disease. In addition, Unicapsula seriolae is also suspected to be a parasite associated with an unidentified foodborne disease of caused by ingestion of Greater amberjack, Seriola dumerili [37]. Screening for K. septempumnctata is carried out on cultured olive flounder by the Korean government, however other myxosporeans containing other Kudoa spp. that infect the fish are not investigated. Because Kudoa spp. have a low host specificity [38], there is a possibility that cultured olive flounders are infected by other Kudoa spp. (or myxosporeans) related with foodborne illness. In addition, most consumers do not want to eat fish infected by any parasites. Thus, more studies investigating other myxosporeans (especially parasites that infect muscle) and developing control methods against parasite infections should be conducted.
ACKNOWLEDGMENTSThis work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2016R1D1A1B03932598) and is part of the project titled ‘Fish Vaccine Research Center’, funded by the Ministry of Oceans and Fisheries, Korea.
Conflict of interestCONFLICT OF INTEREST
The authors declare no conflict of interest related to this study.
REFERENCES1. Chang ES, Neuhof M, Rubinstein ND, Diamant A, Philippe H, Huchon D, Cartwright P. Genomic insights into the evolutionary origin of Myxozoa within Cnidaria. Proc Natl Acad Sci USA 2015;112:14912-14917.
2. Whipps CM, Grossel G, Adlard RD, Yokoyama H, Bryant MS, Munday BL, Kent ML. Phylogeny of the multivalvulidae (Myxozoa: Myxosporea) based on comparative ribosomal DNA sequence analysis. J Parasitol 2004;90:618-622.
3. Eiras JC, Saraiva A, Cruz C. Synopsis of the species of Kudoa Meglitsch, 1947 (Myxozoa: Myxosporea: Multivalvulida). Syst Parasitol 2014;87:153-180.
4. Yokoyama H, Whipps CM, Kent ML, Mizuno K, Kawakami H.
Kudoa thyrsites from Japanese flounder and Kudoa lateolabracis n. sp. from Chinese sea bass: causative myxozoans of post-mortem myoliquefaction. Fish Pathol 2004;39:79-85.
5. Yokoyama H, Suzuki J, Shirakashi S.
Kudoa hexapunctata n. sp. (Myxozoa: Multivalvulida) from the somatic muscle of Pacific bluefin tuna Thunnus orientalis and re-description of K. neothunni in yellow fin tuna T. albacares
. Parasitol Int 2014;63:571-579.
6. Arai Y, Matsumoto K. On a new sporozoa, Hexacapsula neothunni gen. et sp. nov. from the muscle of Yellowfin tuna, Neothunnus macropterus
. Nippon Suisan Gakk 1953;18:293-298.
7. Gilchrist JDF. A protozoal parasite (Chloromyxum thyrsites, sp. n.) of the Cape sea-fish, the “Snoek” (Thyrsites atun, Euphr.). Trans R Soc S Afr 1923;11:263-273.
8. Egusa S, Shiomitsu T. Two new species of the genus Kudoa (Myxosporea: Multivalvulida) from marine cultured fishes in Japan. Fish Pathol 1983;18:163-171.
9. Egusa S, Nakajima K.
Kudoa amamiensis n. sp. (Myxosporea: Multivalvulida) found in cultured Yellowtails and wild Damselfishes from amami-ohshima and okinawa, Japan. Nippon Suisan Gakk 1980;46:1193-1198.
10. Yasunaga N, Hatai K, Ogawa S, Yasumoto S. An unknown Myxozoa found in brain of cultured Sea bass, Lateolabrax japonicus and cultured Japanese striped knifejaw, Oplegnathus fasciatus
. Fish Pathol 1981;16:51-54.
11. Hsieh SE, Chen CL.
Septemcapsula yasunagai gen. et. sp. nov., representative of a new family of the class myxosporea. Acta Zootaxon Sin 1984;9:225-227.
12. Kawai T, Sekizuka T, Yahata Y, Kuroda M, Kumeda Y, Iijima Y, Kamata Y, Sugita-Konishi Y, Ohnishi T. Identification of Kudoa septempunctata as the causative agent of novel food poisoning outbreaks in Japan by consumption of Paralichthys olivaceus in raw fish. Clin Infect Dis 2012;54:1046-1052.
13. Kim JJ, Ryu S, Lee H. Foodborne illness outbreaks in Gyeonggi province, Korea, following seafood consumption potentially caused by Kudoa septempunctata between 2015 and 2016. Osong Public Health Res Perspect 2018;9:66-72.
14. Ohnishi T, Tomaru A, Yoshinari T, Kamata Y, Sugita-Konishi Y. Detection of myxosporean parasites in unidentified food-borne disease associated with consumption of raw fish. Jap J Food Microbiol 2016;33:150-154 (in Japanese)..
15. Shin SP, Sohn HC, Jin CN, Kang BJ, Lee J. Molecular diagnostics for verifying an etiological agent of emaciation disease in cultured olive flounder Paralichthys olivaceus in Korea. Aquaculture 2018;493:18-25.
16. Shin SP, Nam Jin C, Chang Sohn H, Lee J.
Parvicapsula curvatura n. sp. in cultured olive flounder Paralichthys olivaceus and phylogenetic characteristics of the genus Parvicapsula
. Dis Aquat Organ 2018;130:199-207.
17. Shin SP, Jin CN, Sohn HC, Lee J.
Sinuolinea capsularis (Myxosporea: Sinuolineidae) isolated from urinary bladder of cultured olive flounder Paralichthys olivaceus
. Korean J Parasitol 2019;57:127-134.
18. Song JY, Kim MJ, Choi HS, Jung SH. Monitoring Kudoa septempunctata in cultured olive flounder Paralichthys olivaceus in different regions of Korea in 2013. Korean J Fish Aquat Sci 2014;47:611-621 (in Korean)..
19. Cho JB, Kim KH. Light- and electron-microscope description of Kudoa paralichthys n. sp. (Myxozoa, Myxosporea) from the brain of cultured olive flounder Paralichthys olivaceus in Korea. Dis Aquat Organ 2003;55:59-63.
20. Lom J, Arthur JR. A guideline for the preparation of species descriptions in Myxosporea. J Fish Dis 1989;12:151-156.
21. Hillis DM, Dixon MT. Ribosomal DNA: molecular evolution and phylogenetic inference. Q Rev Biol 1991;66:411-453.
22. Fiala I. The phylogeny of Myxosporea (Myxozoa) based on small subunit ribosomal RNA gene analysis. Int J Parasitol 2006;36:1521-1534.
23. Barta JR, Martin DS, Liberator PA, Dashkevicz M, Anderson JW, Feighner SD, Elbrecht A, Perkins-Barrow A, Jenkins MC, Danforth HD, Ruff MD, Profous-Juchelka H. Phylogenetic relationships among eight Eimeria species infecting domestic fowl inferred using complete small subunit ribosomal DNA sequences. J Parasitol 1997;83:262-271.
24. Bartosová P, Fiala I, Hypsa V. Concatenated SSU and LSU rDNA data confirm the main evolutionary trends within myxosporeans (Myxozoa: Myxosporea) and provide an effective tool for their molecular phylogenetics. Mol Phylogenet Evol 2009;53:81-93.
25. Van der Auwera G, Chapelle S, De Wachter R. Structure of the large ribosomal subunit RNA of Phytophthora megasperma, and phylogeny of the oomycetes. FEBS lett 1994;338:133-136.
26. Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG. Clustal W and Clustal X version 2.0. Bioinformatics 2007;23:2947-2948.
27. Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 2016;33:1870-1874.
28. Shin SP, Ishitani H, Shirakashi S. Development of a multiplex PCR to detect Kudoa spp. and to distinguish Kudoa septempunctata in olive flounder Paralichthys olivaceus
. Aquaculture 2016;464:37-41.
29. Matsukane Y, Sato H, Tanaka S, Kamata Y, Sugita-Konishi Y.
Kudoa septempunctata n. sp. (Myxosporea: Multivalvulida) from an aquacultured olive flounder (Paralichthys olivaceus) imported from Korea. Parasitol Res 2010;107:865-872.
30. Kasai A, Li YC, Mafie E, Sato H. New host records of monacanthid fish for three Kudoa spp. (K. septempunctata, K. thyrsites, and K. shiomitsui) prevalent in the olive flounder (Paralichthys olivaceus), with the description of K. parathyrsites n. sp. from a black scraper (Thamnaconus modestus). Parasitol Res 2016;115:2741-2755.
31. Ohnishi T, Kikuchi Y, Furusawa H, Kamata Y, Sugita-Konishi Y.
Kudoa septempunctata invasion increases the permeability of human intestinal epithelial monolayer. Foodborne Pathog Dis 2013;10:137-142.
32. Iwashita Y, Kamijo Y, Nakahashi S, Shindo A, Yokoyama K, Yamamoto A, Omori Y, Ishikura K, Fujioka M, Hatada T, Takeda T, Maruyama K, Imai H. Food poisoning associated with Kudoa septempunctata
. J Emerg Med 2013;44:943-945.
33. Jang Y, Ahn M, Bang H, Kang B. Effects of Kudoa septempunctata genotype ST3 isolate from Korea on ddY suckling mice. Parasite 2016;23:18.
34. Ahn M, Woo H, Kang B, Jang Y, Shin T. Effect of oral administration of Kudoa septempunctata genotype ST3 in adult BALB/c mice. Parasite 2015;22:35.
35. Ahn M, Ko HJ, Kim J, Jang Y, Shin T. Evaluation of the inflammatory response to Kudoa septempunctata genotype ST3 isolated from olive flounder (Paralichthys olivaceus) in Caco-2 cells. Parasite 2018;25:12.
36. Kawase M, Yoshioka A, Hosoya M, Shichiku M. Suspected foodborne disease related to juvenile Pacific bluefin tuna containing Kudoa hexapunctata and evaluation of patient fecal examination. Jap J Food Microbiol 2015;32:48-53 (in Japanese)..
Fig. 1
Kudoa ogawai found in the muscle of Paralichthys olivaceus. (A) Cyst and (B) myxospores of K. ogawai. Scale bar=100 and 20 μm, respectively. (C) Myxospores of K. septempunctata. Scale bar=20 μm. (D) Fresh spores of K. ogawai in apical view and (E) side view. Arrows and arrowheads show the polar capsule and the apical projections of spores, respectively. Scale bar=10 μm. 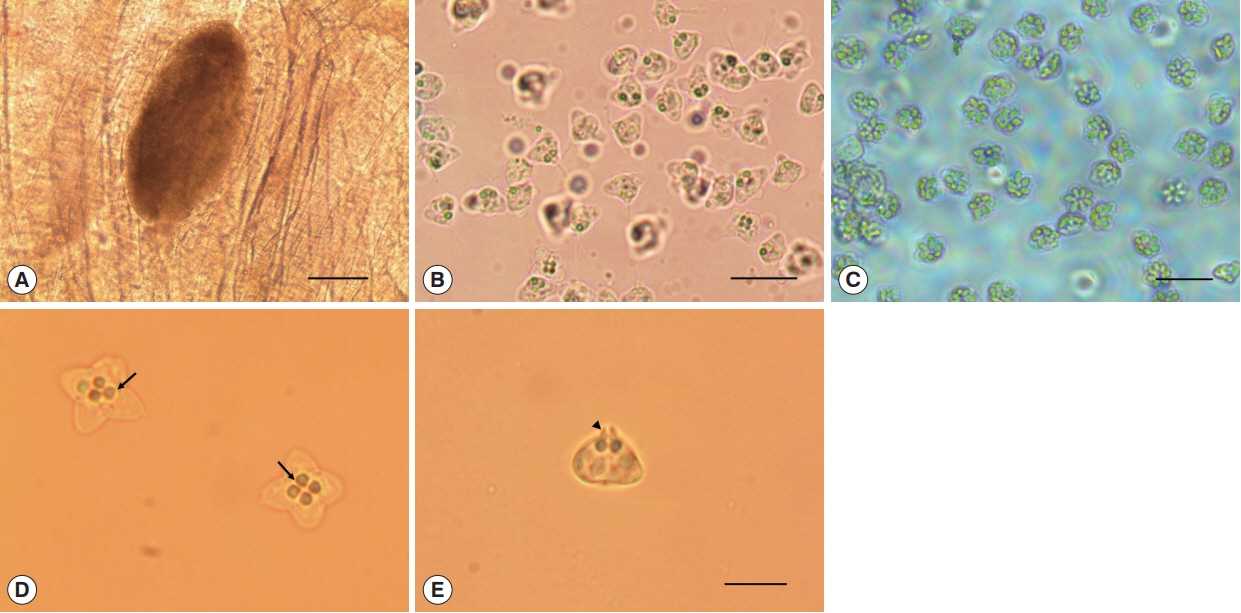
Fig. 2Phylogenetic tree generated by MEGA 7, based on the aligned partial 18S rDNA sequences of Kudoa sp. newly obtained from the present study and Kudoa spp. previously reported in olive flounder. Enteromyxum leei was set as the outgroup and the bootstrap values (1,000 replicates) are shown at the nodes on the neighbor-joining tree. The present species is indicated by an arrow and is clustering with K. ogawai. 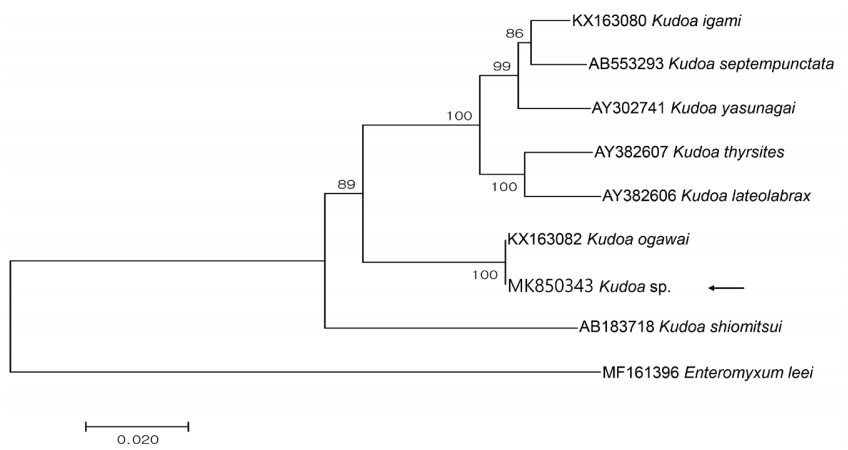
Table 1Comparison of Kudoa sp. (present isolate) with the other Kudoa spp. reported in olive flounders Paralichthys olivaceus
a Data provided from K. lateolabracis isolated from Lateolabrax sp. at the original description (Yokoyama et al., 2004 [4]). b Data provided from K. yasunagai isolated from Lateolabrax japonicus (Hsieh et al., 1984 [11]). c Data provided from K. shiomitsui isolated from Stephanolepis cirrhifer (Kasai et al., 2016 [30]). |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||