INTRODUCTION
Clonorchis sinensis is a fish-borne trematode causing an endemic disease in East Asian countries such as Korea, China, and northern Vietnam. About 35 million people are estimated to be infected with this trematode [1]. The adult flukes inhabit in the intrahepatic bile ducts of the fish-eating mammals. When the mammals consume infected freshwater fish, the metacercariae excyst in the duodenum of these mammals. The excysted larvae migrate through the ampulla of Vater and common bile duct and finally reach the intrahepatic biliary duct. Adult worms commonly parasitize in the intrahepatic ducts and provoke pathologic changes in the biliary tracts. This fluke provokes, in some cases, cholangiocarcinoma [2-4].
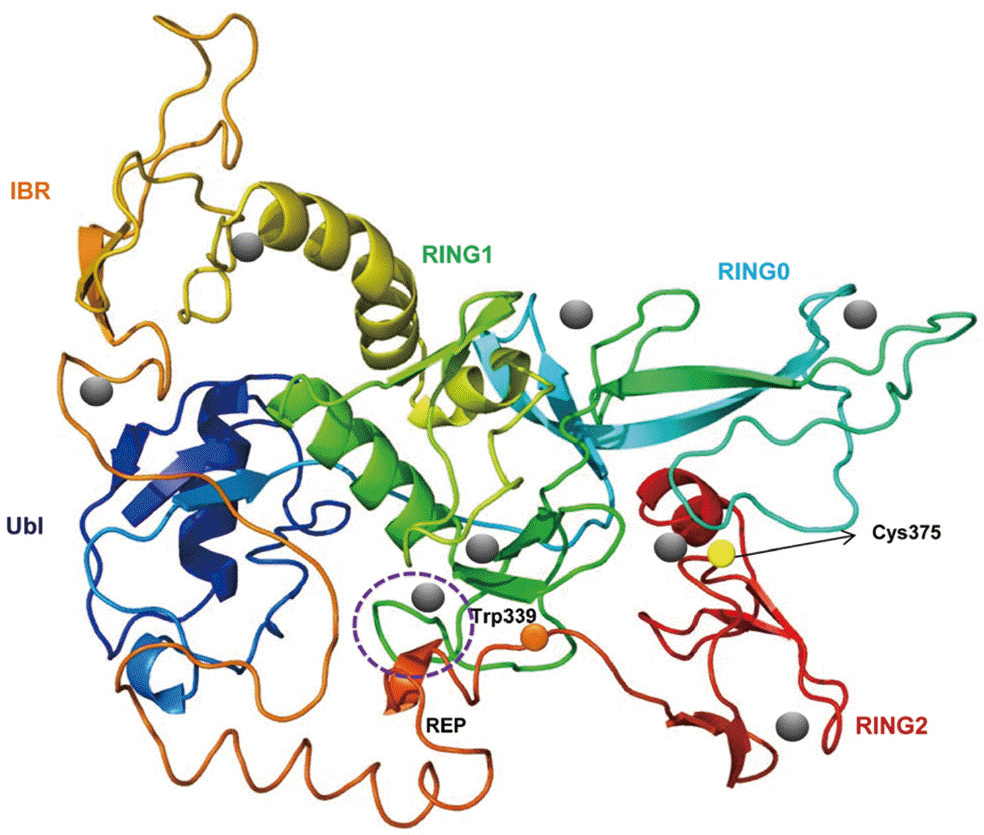
From the C. sinensis transcriptome and genome, many genes were characterized to play important roles in C. sinensis metabolism and/or pathogenesis to the hosts, but not parkin gene yet. Parkin is a causative factor of autosomal recessive juvenile Parkinsonism characterized by the loss of dopaminergic neurons in substantia nigra with symptoms such as tremor, rigidity, and bradykinesia [5]. The 465-residue human parkin is a RING-between-RING (RBR) protein consisting of 5 domains: N-terminal ubiquitin-like domain (Ubl), 3 RING domains, and an in-between-ring domain (IBR). The RING domains have each 8 cysteine/histidine residues in the general formula C-x2-C-x(9-39)-C-x(1-3)-H-x(2-3)-C-x2-C-x(4-48)-C-x2-C, which bind 2 Zn2+ ions and form a cross-brace, which does not resemble the classical zinc fingers [6]. The IBR domain connects the RING1 and RING2 domains [7-9].
Parkin has been identified in diverse cellular processes using substrates such as α-synuclein [10], synphilin-1 [11], Sept5 [12], Pael-R [13], cyclin E [14], α/β tubulin [15], and JTV1 [16], suggesting indirect transcriptional and metabolic regulations. Parkin possesses ubiquitin ligase activity that catalyzes transfer of ubiquitin conjugated with E2 to specific protein substrates [17]. This is an important post-translational modification process of proteins. The ubiquitinated proteins are processed either to proteolytic degradation or to intracellular location. Parkin regulates internalization and degradation of epidermal growth factor receptor (EGFR) through ubiquitination of Eps15 [18]. Parkin ubiquitinates TDP-43 and facilitates its cytosolic accumulation [19]. Dysfunction of parkin results in accumulation of its substrate which may lead to unfolded protein stress and cell death. In addition, parkin reduces oxidative stress in mitochondria and maintains homeostasis [20,21]. In the genome databases, the parkin genes identified are of humans, rats, mice, and Drosophila [22,23].
This study was carried out to identify C. sinensis parkin (CsParkin) on a cDNA clone retrieved from the C. sinensis transcriptome and to analyze its molecular structures and biological characteristics.
MATERIALS AND METHODS
Ethics statement
BALB/c mice (female, 7-week-old) and rabbits (New Zealand White, male, 2.2-2.5 kg) were handled in an accredited Chung-Ang University animal facility (accredited unit, Korea FDA; unit no. 36). Approval for animal experiments was obtained from Institutional Animal Care and Use Committee of Chung-Ang University (permission no. CAU-2011-0013). This study was carried out strictly in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the United States National Institutes of Health. Freshwater fish Pseudorasbora parva were purchased from a city market in Shenyang, Liaoning Province, P. R. China. This activity did not require a specific permission, as it did not involve any endangered or protected species in the field.
Bioinformatics analysis, structure modeling, and phylogenetic tree construction
An EST clone encoding a polypeptide homologous with parkin of other animals was found through analysis on the C. sinensis transcriptome database using bioinformatics tools. This EST clone (CSA08099) was retrieved from C. sinensis transcriptome glycerol stock [24] and was sequenced further to reach poly (A) tail. On this cDNA sequence, putative polypeptide sequence with open reading frame was predicted using ExPASy translate tool (http://web.expasy.org/translate/). BLASTx tool was used to search CsParkin homologues in the NCBI database. CsParkin polypeptide was aligned with parkin of other animals using Clustal X and curated manually in GeneDoc [25], then functional domains of CsParkin were identified. The secondary and tertiary structures of CsParkin were predicted on a template rat parkin [26] using Phyre2 [27]. A phylogenetic tree was drawn with 24 parkins retrieved from GenBank using Neighbor-Joining algorhism in Mega 5.2 software [28]. Bootstrap value was of 1,000 replicates.
Production of recombinant CsParkin
The coding region of CsParkin cDNA was amplified by PCR using specific forward (5´-CAGATACGGAATTCATGCTTGTACGGATCTCTCAC-3´) and reverse (5´-CAGATAGT CTCGAGTTAGAGCCAATGTAAGGCTTG-3´) primers containing EcoRI or XhoI restriction enzyme site. Amplification was started with denaturing DNA at 95˚C for 5 min then followed by amplification phase of 30 thermal cycles (95˚C for 1 min, 65˚C for 30 sec, 72˚C for 2 min) and a final extension at 72˚C for 10 minPCR product was purified using a spin column (QIAGEN, Hilden, Germany), digested with the restriction enzymes and subcloned into an expression plasmid vector pGEX-4T-1. In this expression construct, recombinant CsPArkin is produced as a fusion protein to a tag protein Sj26GST. Escherichia coli BL21 (DE3) pLysS (Invitrogen, Carlsbad, California, USA) was transformed with the expression plasmid DNA. The transformed E. coli were liquid-cultured overnight at 18˚C and recombinant CsParkin protein was induced by adding isopropyl β-D-thiogalactopyranoside (IPTG) at a final concentration of 0.2 mM with supplementation of 1 mM ZnCl2. E. coli was harvested by spinning and homogenized in 1×PBS by sonication at 1 sec intervals for 10 min (35 vibrations/pulse). After spinning, the supernatant was saved and loaded onto a glutathione sepharose 4B column (GE Healthcare, Uppsala, Sweden) at 4˚C for 3 hr. Recombinant CsParkin was cleaved from sj26GST tag protein by thrombin treatment overnight at 4˚C and eluted with 1× PBS. The recombinant CsParkin protein was subjected to 10% SDS-PAGE. Protein concentration was determined using a protein assay kit (Bio-Rad, Hercules, California, USA).
Production of partial CsParkin
To produce sufficient amount of protein for mouse immunization, recombinant CsParkin protein was produced in a partial form in E. coli. B cell epitopes were predicted on the CsParkin polypeptide sequence using IEDB program (http://tools.immuneepitope.org/tools/bcell/iedb_input). A fragment of high immunogenicity (estimated molecular mass 21 kDa) was selected and its coding cDNA region was PCR-amplified using specific forward (5’- GACTCGGAATTCTTGCAACTAAGGTTCGATG-3’) and reverse (5’-AGATCTCGAG TCCACTCTCCAAGTACTCC-3’) primers containing EcoRI or XhoI restriction enzyme site. Amplicon was purified and digested with the restriction enzymes and was subcloned into pET-23a (+). After transforming E. coli BL21 (DE3) pLysS, the partial recombinant protein was induced using 0.1 mM IPTG for 4 hr at 37˚C. E. coli cells were harvested and lyzed in 8M urea by sonication. After spinning, supernatant containing partial CsParkin protein was loaded on Ni-NTA agarose (QIAGEN) column at a denaturing condition and stood at room temperature for 3 hr. The Ni-NTA agarose column was washed twice with 8M urea buffer and recombinant protein was eluted with 8M urea buffer containing 100 mM imidazole. The partial recombinant CsParkin protein was dialyzed against 1x PBS and checked for quality by 10% SDS-PAGE.
Production of immune sera
Female, 7-week-old BALB/c, mice were purchased from an animal supply company (Samtako BioKorea Co., Osan, Korea). For primary immunization, recombinant partial CsParkin protein (50 μg per mouse) was emulsified with Freund’s complete adjuvant and injected into the mouse peritoneum. Two weeks later, mice were injected intraperitoneally with a booster dose of 50 μg recombinant protein emulsified with Freund’s incomplete adjuvant. Blood was taken from eye fundus of the immunized mice 2 weeks after the booster injection and anti-CsParkin sera were separated from the blood.
Metacercariae and adults of C. sinensis
Freshwater fish, P. parva, the second intermediate host of C. sinensis, were digested with artificial gastric juice (0.8% pepsin, 0.8% HCl). After sedimenting and washing, C. sinensis metacercariae were collected from sediments under a dissection microscope. The metacercariae were fed twice intragastrically to rabbits with 1 week interval. Six weeks after the second infection, C. sinensis adult worms were recovered from the rabbits’ bile duct and stored at -80˚C in 1× PBS until used.
Western blotting of native CsParkin
C. sinensis adults were homogenized in 2 volumes of 1× PBS containing 1% Triton X-100 and 1× Complete™ mini (Roche, Mannheim, Germany) and kept at 4˚C overnight. The suspension was centrifuged at 13,000 rpm for 60 min at 4˚C. The second supernatant was used as a crude extract of C. sinensis adults. The recombinant partial CsParkin protein and the C. sinensis adult extract were resolved by 10% SDS-PAGE and transferred onto a nitrocellulose membrane. The membrane was blocked within 5% skim milk at room temperature for 1 hr and incubated in anti-CsParkin mouse serum at dilution of 1:100 at 4˚C overnight. After washing 3 times with PBS-Tween 20, the membrane was incubated with goat anti-mouse IgG alkaline phosphatase-conjugated antibody (Sigma-Aldrich, St. Louis, Missouri, USA) at dilution of 1:5,000 at room temperature for 2 hr. Color was visualized using a substrate BCIP/NBT (Si
gma-Aldrich).
C. sinensis total RNA and quantitative real-time PCR
C. sinensis adult and metacercariae were ground into powder in liquid nitrogen using mortar and pestle. Total RNA was extracted using TRIzol reagent (Ambion, Carlsbad, California, USA) according to the manufacturer’s instruction. From the total RNA, trace of genomic DNA was removed using DNA-free kit (Ambion). Quality of the total RNA was evaluated by determining the OD260/280 using a spectrophotometer. The first-strand cDNA was synthesized using power cDNA synthesis kit with oligo-d(T) primer (INTRON Biotechnology, Gyeonggi-do, Korea) according to the manusfacturer’s protocol.
To perform quantitative real time-PCR (qRT-PCR), primers for CsParkin were designed using the Oligo ver. 6.71 software (Molecular Biology Insights, Cascade, Colorado, USA). Three genes, i.e., β-actin, calcyphosine and phosphoglycerate kinase, expressed stably between the adult and the metacercaria [29], were employed as reference genes. The primer sequences used for qRT-PCR are in Table 1. Reaction mix of 10 μl was made of 70 ng total cDNA of the adults or metacercariae, primer sets each of CsParkin, β-actin, calcyphosine or phosphoglycerate kinase and 1x master mix (FasStart SYBR Green I Kit, Roche). Thermal cycle was performed using LightCycler 1.5 (Roche) as follows: the reaction mix was heated to 95˚C for 15 min, followed by 45 cycles of 95˚C for 10 sec, 60˚C for 10 sec and 72˚C for 30 sec. A developmental expression of CsParkin was calculated using 2-ΔΔCt method and presented as fold-difference [30].
Ubiquitination assay in vitro
Ubiquitination assay was performed in vitro using Parkin Auto-Ubiquitination kit (BostonBiochem, Cambridge, Massachusetts, USA). The full-length recombinant CsParkin was incubated with 1 mM Mg-ATP, 50 mM HEPES buffer, pH 7.5, 50 mM NaCl, 1 mM dithiothreitol, 11 μg/ml of human E1, 9 μg/ml of human UbcH7, and 0.1 mM of human ubiquitin at 37˚C for 1 hr. For positive control reaction, human parkin was employed as a reference from the kit. After incubation, the reaction products were deployed by 10% SDS-PAGE and transferred onto a nitrocellulose membrane. The membrane was immunoblotted using mouse anti-human ubiquitin IgG (BostonBiochem) and alkaline phosphatase-conjugated goat anti-mouse IgG antibody (Sigma-Aldrich).
Immunohistochemical staining of CsParkin in C. sinensis adults and metacercariae
C. sinensis adults in the rabbit liver and metacercariae from fish were fixed in 10% paraformaldehyde, embedded in paraffin, and sectioned into ribbons. The ribbons were deparaffinized, rehydrated, and treated with anti-CsParkin mouse sera or naive mouse sera at a dilution of 1:50 for the adult flukes or a 1:100 for the metacercariae for 2 hr at room temperature. Subsequently, the ribbons were incubated with polymer-horse radish peroxidase labeled anti-mouse IgG antibody (Dako Cytomation, Glostrup, Denmark). Color was developed using 3-amino-9-ethylcarbazole as substrate and the ribbons were counterstained with hematoxylin.
RESULTS
Molecular structure and phylogeny of CsParkin
The CsParkin cDNA was 1,218 nucleotides long and encoded a putative polypeptide of 405 amino acids with an estimated molecular weight of 45.7 kDa. In the polypeptide, highly conserved were Cys and His residues which bind to Zn2+ ion and form RING domains (Fig. 1). Secondary structure of CsParkin revealed 5 domains in tandem: ubiquitin-like (Ubl), RING0, RING1, in-between-RING (IBR), and RING2. CsParkin lacked of a long tether between the Ubl and RING0 domains, but found in other animals. In Ubl domain, a hydrophobic RING1-binding site was conserved at Leu43. RING1 was a canonical RING finger with classic cross-brace arrangement and had an E2-binding site consisting of Ala164, Phe166, and Asn167. In all parkins, a linker between IBR and RING2, termed a repressor element of parkin (REP) [26], displayed low similarity, but a residue Trp339 was highly conserved. A catalytic motif consisting of Cys375, X376, and His377 was well conserved and evident in CsParkin RING2 domain (Fig. 2).
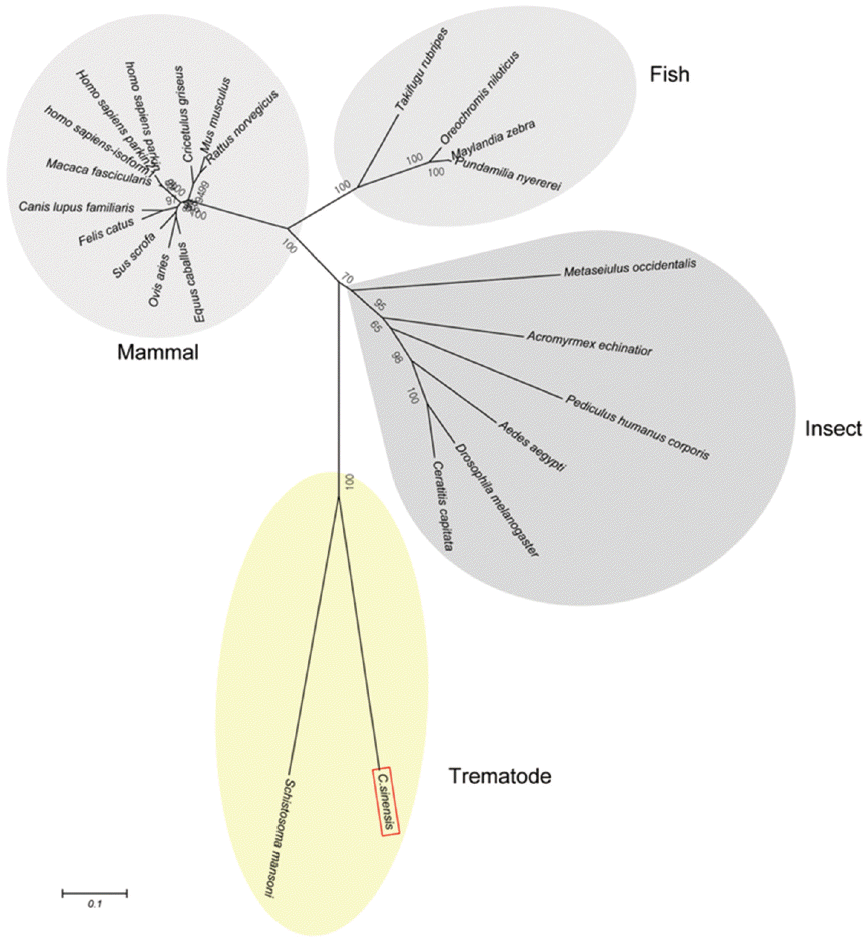
A predicted tertiary structure of CsParkin showed a very similar conformation to the crystal structure of the parkin of human and rat [26,31]. IBR was conformationally adjacent to RING1, while RING0 was adjacent to RING2. The E2-binding site in RING1 was occupied by REP near the Ubl binding site (Fig. 2). A phylogenetic tree showed CsParkin grouped with a homolog of blood fluke, Schistosoma mansoni. Parkin has not been reported from other trematodes and Platyhelminthes yet. Parkins of the mammalian animals formed a branch appearing as a major clade, followed by clades of fish and the insects (Fig. 3).
Recombinant and native CsParkin
When the recombinant full-length CsParkin fused with Sj26GST tag protein (70 kDa as whole molecule) was overexpressed in E. coli BL21 (DE3) pLysS, it remained insoluble in E. coli homogenate. Adding Zn2+ to liuid culture medium alleviate this odd and resolved a small portion of the recombinant fusion protein to soluble in E. coli. This soluble fusion protein bound efficiently to GSH-Sepharose beads. By thrombin digestion, the recombinant full-length CsParkin was on-bead cleaved from tag-protein Sj26GST and eluted as a single fraction to homogeneity. When dialyzed against PBS, part of the recombinant CsParkin remained in soluble part, but some turned to insoluble and sedimented as aggregate. This soluble CsParkin was used for its functional assay, ligase activity.
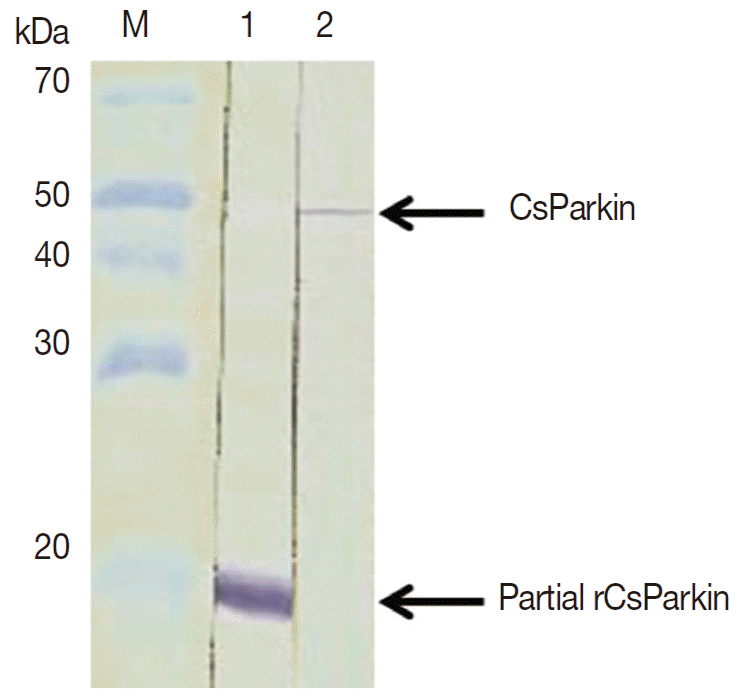
To produce a soluble form, a partial CsParkin, 21 kDa, with 6×His tag was produced in E. coli BL21 (DE3) pLysS and purified using Ni-NTA agarose column under denaturing condition. After dialyzing against 1x PBS, the purified recombinant partial CsParkin protein remained totally in soluble fraction and near to homogeneity. This partial CsParkin was used for the production of mouse immune sera. The mouse immune sera recognized a native CsParkin showing a molecular mass close to estimated 45.7 kDa (Fig. 4).
Relative expression of CsParkin in the developmental stages
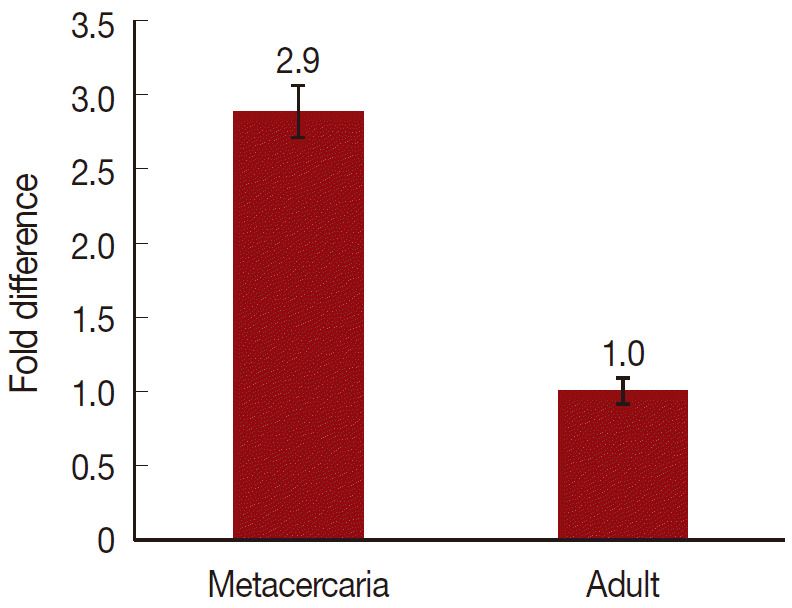
On a total cDNA as template, developmental expresion of CsParkin in the C. sinensis adults and metacercariae was determined by qRT-PCR. CsParkin was expressed 2.9 folds higher in the metacercariae than in the adults (Fig. 5).
In vitro auto-ubiquitination of CsParkin
To demonstrate ubiquitin ligase activity of CsParkin, an in vitro ubiquitination assay was performed with the full-length recombinant CsParkin. In the in vitro auto-ubiquitination assay, compared to a strong band in the positive control, a small band appeared in the full-length recombinant CsParkin lane, implying that auto-ubiquitination with a heterologous human ubiquitin was taken place (Fig. 6).
Distribution of CsParkin
Immunohistochemical staining using mouse immune sera revealed native CsParkin in the developmental stages. In the adult worms, CsParkin was localized in the testes, sperms in the seminal receptacle and oral sucker. Weak staining was recognized in the mesenchymal tissues. In metacercariae, the CsParkin was extensively localized in the tegument, subtegument, mesenchymal tissues, and oral and ventral suckers (Fig. 7).
DISCUSSION
Parkin has been cloned from several animals and its diverse functions have been characterized. In this study, a parkin homolog was identified from C. sinensis and characterized for its molecular and biological functions. The putative polypeptide of CsParkin showed a high conservation of Cys and His residues, involving 3 RING domains aligned in a R0-RBR structure. The Ubl domain could bind to the S5a subunit of human 26S proteasomes or to ubiquitin-interacting motifs (UIMs) in a substrate. In both interactions, Arg42 in the Ubl domain plays a crucial role [18,32]. This positively charged Arg42 residue was replaced with negatively charged Asp41 in CsParkin or with Glu49 in Caenorhabditis elegans parkin. Whether these residue replacements influence molecular function is unclear. The Ubl domain binds conformationally to the E2-binding site in the RING1 domain; this interaction was recently found in the rat parkin through crystalography [26]. This conformation inhibits auto-ubiquitination of parkin [33]. The RING1-binding site in Ubl domain of CsParkin included hydrophobic Leu42 and Leu43, very similar with other parkin homologs, suggesting an inhibitory function of CsParkin. A short tether loop between the Ubl and RING0 domains in the parkin of lower animals such as C. sinensis and C. elegans may allow less flexibility to tertiary structure and influence functionality.
CsParkin RING0 domain had an array of conserved (C4C3 H), as in vertebrate animals, and was predicted to coordinate with 2 Zn2+ ions. With high similarity to human parkin, the RING1 domain of CsParkin is a canonical RING finger with classic cross-brace arrangement of Cys and His residues. In the E2-conjugating enzyme binding site in the RING1 domain of CsParkin, Ala164 was highly conserved and 2 residues were substituted with Phe166 and Asn167, suggesting that CsParkin could mediate ubiquitin transfer. The IBR domain forms a pair of scissors-like and GAG knuckle-like zinc-binding sites, which bring 2 RING fingers close and facilitate protein interactions. In IBR of CsParkin, even though Cys residues were spaced longer than in those of homologues, they could flexibly bind to Zn2+ and form an IBR structure. Eventhough the REP linker between IBR and RING2 domains is dissimilar among all parkins, with one highly conserved Trp339, it plays a crucial role in repressing parkin ligase activity. Among the 3 RING domains, the RING2 domain shows the highest homology in all parkins, indicating that this region is evolutionarily conserved among all animals. As CsParkin had an E2-binding site in RING1 and catalytic residues conserved in RING2, it is suggested that CsParkin could have E3 ligase activity. The E3 ligase has dual activities of RING- and HECT-type ligases; first transfers ubiquitin from the E2 to the RING2 domain of the E3 and then to the substrate. For correct folding and structural stability of parkin proteins, Zn2+-binding to R0-RBR domains are indispensable. In producing recombinant CsParkin in E. coli, supplementation of the culture medium with Zn2+ may facilitate correct folding of the recombinant protein and retain its solubility.
The best described function of the parkin is E3 ligase activity, which interferes with ubiquitin-mediated proteolytic pathway. For CsParkin, auto-ubiquitination was detected in vitro ubiquitination assay. E3 ligase activity of CsParkin was much weaker than that of human parkin since the partner proteins added to the reaction system were heterologous, not of C. sinensis origin, which could attenuate binding of CsParkin to the substrates.
In recent years, studies have been focused on further roles of parkin in mitochondrial functionality. As an energy conversion and supplying organelle, mitochondrial homeostasis is essential to maintain cellular normality. Parkin is recruited from cytoplasm to damaged mitochondria and mediates proteasome-dependent degradation of inner and outer membrane [34,35]. Parkin also regulates mitochondrial spheroid formation and mitophagy via degradation of mitofusins. Mitochondrial hexokinase HK1, a pivotal enzyme involved in the cellular uptake and utilization of glucose, is a substrate of parkin, implying that parkin regulates cellular energy metabolism [36]. Decrease in parkin expression leads to mitochondrial DNA damage and morphological change. Deficiency of parkin provokes mitochondrial dysfunction, decrease of antioxidant capacity and proportional change of proteins associated with energy metabolism [20,37]. In this study, CsParkin was found mainly distributed in the reproductive and locomotive organs of the C. sinensis adults. As the parasites growing, the C. sinensis adults produce a lot of eggs, which require large amount of energy and metabolic intermediates produced from glucose [38,39]. The oral sucker is a main locomotorium of C. sinensis, reaching out and holding fast at a point then tracking the hind body. For this activity, the oral sucker, which is composed of many myocytes, consumes a great deal of energy supplied by mitochondria [39,40]. In the Parkin-null Drosophila, severe disruption of mitochondria results in apoptotic muscle cell death and deforming of sperms [41]. In these considerations, it is suggested that CsParkin plays a role in mitochondrial energy metabolism.
CsParkin was expressed relatively higher levels and distributed more extensively in the metacercariae than in the adults of C. sinensis. The metacercaria is a larval and resting stage with basal energy metabolism. The metacercariae has exclusively somatic organs and a genital anlage. The anlage appears as an indistinctive primodium posterior to the ventral sucker and differentiate into reproductive organs during development to adult C. sinensis. Parkin regulates phospholipase C-γ1 and arrestin in the cell differentiation and signaling pathways [42,43]. Parkin can regulate EGFR through ubiquitination of Eps15, but does not bind to EFGR. Transcription levels of EGF and TGF-beta interacting protein 1 were higher in the metacercaria than in the adult C. sinensis [24]. It is suggested that CsParkin regulate expression of some genes, which are related to cellular proliferation and differentiation and signaling pathways in the juvenile C. sinensis developing to the adult.