Blastocystis is a genus of intestinal parasites detected in humans and other animals worldwide and is typically found as cysts in feces [1]. The pathway of infection is thought to involve the ingestion of water or food containing cysts spread from the feces of infected animals. The prevalence has been reported to be higher in developing countries than in developed countries because of the less hygienic conditions in the former [2]. The clinical symptoms of Blastocystis infection include abdominal pain, diarrhea, flatulence, and fatigue [3]. The infection may be asymptomatic or chronic. Clinical differences may also depend on the genotype differences of Blastocystis [4]. A total of 17 subtypes (STs) of Blastocystis have been classified based on their 18S rRNA sequences in mammals and birds [5]. Blastocystis shows low host specificity but has been reported in a variety of hosts, and host-to-host transmission is possible, indicating the potential for zoonotic transmission [6]. In humans, 9 STs of Blastocystis have been detected (ST1–ST9); the most common STs are ST1–ST4, and the predominant type is ST3, which is present in approximately 60% of cases [7]. Additionally, non-human-specific STs have been detected in other animals. Particularly, STs infecting animal handlers have been found to be the same as those isolated from animal feces in contact with the handlers [8]. Because these STs are identical, transmission may occur from animals to humans. Accordingly, a study has reported various hosts and genetic diversity based on the zoonotic potential of Blastocystis [7], and the host range and new STs of Blastocystis have been examined in many countries. Blastocystis has been reported as a common intestinal parasite in pigs in several geographical regions worldwide [1,2,9] and thus has been actively investigated in various countries. Studies on the prevalence, risk factors, genetic characteristics, and STs have been conducted to evaluate the zoonotic potential of Blastocystis infection in pigs over the last decade [8–10]. Recently, Iran reported the first detection of ST6 and ST7 in Asia by sequence analysis of the “barcoding region” [11]. In Brazil, genetic diversity analysis revealed a potentially novel ST [12]. In China, new STs have been reported in goats and sheep, and in the UK, non-primate animals were found to be infected with Blastocystis [13,14]. However, no study has examined Blastocystis infection in Korean pigs. Therefore, this study was conducted to determine the genetic diversity and zoonotic potential of Blastocystis by analyzing pig fecal samples with PCR. Furthermore, we statistically analyzed the prevalence and risk factors and assessed the phylogenetic characteristics of Blastocystis spp. in pigs.
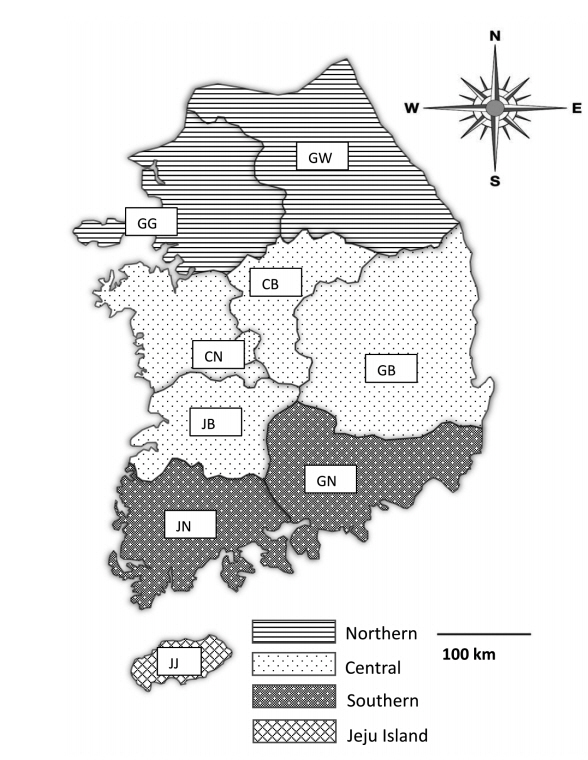
Fecal samples were collected from 28 pig farms rearing more than 1,000 pigs in Korea from May 2017 to August 2019. The farms were chosen only with farm owners’ permission because they were reluctant to allow visitors to enter the farm due to concerns regarding foot and mouth disease, African swine fever, and other infectious diseases. Practicing veterinarians collected fecal samples by rectal palpation. Sample collection did not harm the animals; thus, welfare and ethical approval was not required. The samples were then sent to the Animal and Plant Quarantine Agency (Gimcheon, Korea) and Laboratory of Veterinary Parasitology at Kyungpook National University College of Veterinary Medicine (Daegu, Korea) for parasite examination. The data for each sample, including age, fecal type, collection season, and reared region, were recorded. The pigs were divided into 5 age groups: piglets (<1 month), weaners (<2 months), growers (<4 months), finishers (<6 months), and sows (≥6 months; only females were in this age group). The regions were divided into 4 groups: northern (Gyeonggi [GG] and Gangwon [GW]), central (Chungnam [CN], Chungbuk [CB], Gyeongbuk [GB], and Jeonbuk [JB]), southern (Gyeongnam [GN], Jeonnam [JN]), and Jeju (JJ) (Fig. 1). The minimum number of samples required for the study was 246 according to the formula
n = 1.96 2 p e x p ( 1 - p e x p ) d 2
The statistical significance was determined with the Chi-square test using SPSS version 23.0 software (SPSS Inc., Chicago, Illinois, USA). A P-value of less than 0.05 was considered to indicate significant results. Phylogenetic analysis was performed using the Blastocystis 18S rRNA gene, and a phylogenetic tree was constructed using MEGA 7 software [16]. Phylogenetic inference was conducted using the maximum likelihood method with 1,000 bootstrap replicates.
The PCR results were evaluated according to age, fecal type, season, and region, and the overall prevalence of Blastocystis was 60.4% (Table 1). Most positive samples were detected in the northern region (79.2%) during the winter (78.7%). The infection rate was over 50% in all age groups except the piglet group. The prevalence rates were 39.8%, 60.8%, 77.1%, 64.8%, and 52.3% in the piglet, weaner, grower, finisher, and sow groups, respectively. Fecal type analysis revealed that 61.2% and 59.3% of the DNA samples from diarrhea and normal feces, respectively, were positive.
According to the sequencing results, 80 sequences were divided into 4 STs: ST1, ST2, ST3, and ST5. STs were analyzed according to the previous study [8] and identified by NCBI blast search and phylogenetic analysis. ST distribution revealed that the dominant type was ST5 (70.0%), followed by ST1 (17.5%), ST3 (8.8%), and ST2 (3.8%) (Table 2).
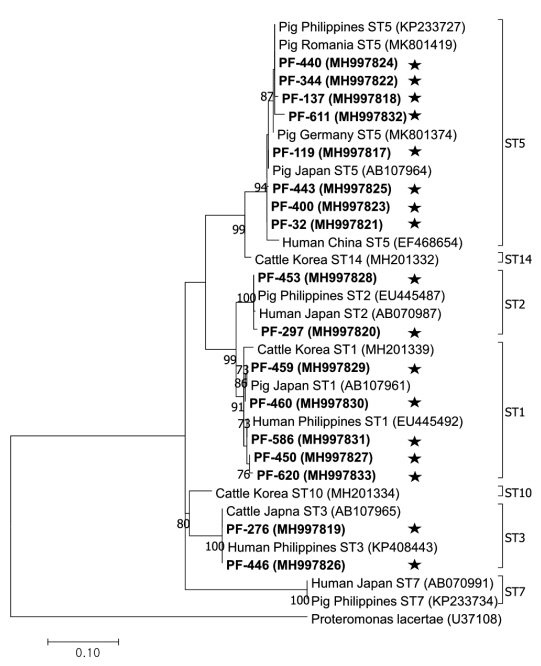
The phylogenetic tree was constructed using 17 sequences representative for each region and 17 GenBank database sequences with Proteromonas lacertae (U37108) as the outgroup (Fig. 2). Each ST detected in this study contained 1 or more closely related Blastocystis sequences identified from humans: 8 sequences belonged to ST5 (MH997817, MH997818, MH997821, MH997822, MH997823, MH997824, MH997825, and MH997832), 5 sequences belonged to ST1 (MH997827, MH997829, MH997830, MH997831, and MH997833), 2 sequences belonged to ST3 (MH997819 and MH997826), and 2 sequences belonged to ST2 (MH997820 and MH997828).
The prevalence of Blastocystis infection has been investigated in various countries worldwide. The infection rate of Blastocystis in pigs was reported as 7.5% and 46.8% in Spain (Aragon and Valencia, respectively), 28.6% in the UK, 34.4% in the USA, 76.7% in Australia, 45.2% in Cambodia, and 100.0% in Vietnam [9]. This genus shows large differences between countries and regions. Previous studies have not evaluated Blastocystis infection in pigs in Korea. Nevertheless, the occurrence of Blastocystis infection in cattle has been examined [17], and Blastocystis infection in pigs in Shanxi province in China has been investigated [9]. However, direct comparisons with Blastocystis infection in Korean pigs are difficult because of differences in the Blastocystis infected animals and geographical and breeding environments. The overall infection rate observed in this study was as high as 60.4%, which may suggest that pigs are natural host [7,8].
Blastocystis infection rates differed significantly according to season (P<0.001). Previous studies revealed high infection rates in hot and rainy seasons [18,19]. However, in this study, the season with the highest prevalence was winter, followed by autumn. These contrasting results may be attributed to differences in the number of samples tested. In the southern region, where the most samples were collected (406/646), more samples were obtained during the autumn and winter (258 samples) than during the spring and summer (148 samples). Therefore, the seasonal infection rates may have been affected by differences in sampling times, limiting the generalization of our results. Regional analysis showed that the infection rate was the highest in the northern region (79.2%), followed by Jeju (63.0%), the central region (58.8%), and the southern region (58.4%). Although the geographic and climatic conditions were not significantly different except for Jeju Island, regional differences in the infection rate were large, which may be attributed to the pig-rearing environment rather than the region. Correlations between sanitation problems and infection rates have been reported previously [20]. In a southern region farm, the management of pigs is controlled by information and communication technology, which regulates the temperature, humidity, and nutrition of the farm. This regulation is thought to be responsible for the low infection rate (35.3%). In some other southern region farms, the positive rates among growers and finishers were high (90.0% and 87.5%, respectively); however, the infection rates were lower in other groups. This may be explained by the rearing of pigs of different ages on separate farms. Additionally, for pig-rearing farms in Korea, grower and finisher pigs are generally bred in large numbers in small spaces, which may increase the chance of contact between individual pigs.
There were significant differences in the infection rates between age groups. Approximately 39.8% and 77.1% of piglets and growers, respectively, were infected. The infection rates in the grower and finisher groups were relatively high, whereas the rate in the piglet group was the lowest. Similar results were reported previously and have been described in relation to the immune system [9,10]. The prevalence of Blastocystis among 4–6-month-old pigs was significantly higher (95.9%) than among pigs of other ages, with the lowest prevalence among 1–2-month-old pigs (57.1%, P<0.05) [9]. Therefore, age is a risk factor in Blastocystis infection. The presence or absence of diarrhea did not appear to significantly affect infection rates; thus, we did not determine the Blastocystis infection status based on this condition. This suggests that Blastocystis infection in pigs does not necessarily lead to diarrhea; similar results were reported for shelter-resident dogs and cats in the USA [21].
In this study, 4 STs (ST1, ST2, ST3, and ST5) were confirmed by NCBI blast search and phylogenetic analysis and found to be closely related to the sequences of STs detected from humans. ST5 was predominant (70.0%) in Korean pigs and a previous study identified this phenomenon in compilation of data in Blastocystis sp. STs [7]. For this reason, ST5 was proposed as pig adapted ST. Other STs including ST1 (17.5%), ST3 (8.8%), and ST2 (3.8%) were detected in this study and similar results were reported in other studies [7,8]. Most pigs contained ST5 or ST1, whereas ST2 and ST3 were rarely detected. However, ST3 is most commonly found in humans [8]. Furthermore, a study has described the potential zoonotic properties of ST5 [8]. In domestic pigs, all of these STs have been detected, demonstrating the genetic diversity of Blastocystis in pigs. Additionally, ST5 was found to be widely distributed throughout Korea. The distribution of all STs in the southern region was confirmed in this study. ST1 and ST3 were detected in the southern and Jeju regions, and ST2 was detected only in the southern region. The largest number of samples was collected in the southern region. Therefore, if the number of samples collected was larger, other STs may be identified in other regions.
In conclusion, the overall Blastocystis infection rate was 60.4% in pigs in Korea, and age was a significant risk factor. There were also regional differences, possibly because of differences in the breeding environment. Sequencing analysis confirmed the presence of 4 genetic Blastocystis STs in Korean pigs, which were also detected in humans, indicating the zoonotic potential of this genus. This is the first study of Blastocystis infection in pigs in Korea. We provide basic information on Blastocystis infection in domestic pig farms. However, limited data were collected for some parts of Korea. Therefore, it is necessary to determine the nationwide infection status and genetic characteristics in further studies. Additionally, as a correlation between the breeding environment and infection rate was observed in this study, further comprehensive studies to evaluate pig farm facilities and environmental management are needed.