Loop-Mediated Isothermal Amplification Targeting Actin DNA of Trichomonas vaginalis
Article information
Abstract
Trichomoniasis caused by Trichomonas vaginalis is a common sexually transmitted disease. Its association with several health problems, including preterm birth, pelvic inflammatory disease, cervical cancer, and transmission of human immunodeficiency virus, emphasizes the importance of improved access to early and accurate detection of T. vaginalis. In this study, a rapid and efficient loop-mediated isothermal amplification-based method for the detection of T. vaginalis was developed and validated, using vaginal swab specimens from subjects suspected to have trichomoniasis. The LAMP assay targeting the actin gene was highly sensitive with detection limits of 1 trichomonad and 1 pg of T. vaginalis DNA per reaction, and specifically amplified the target gene only from T. vaginalis. Validation of this assay showed that it had the highest sensitivity and better agreement with PCR (used as the gold standard) compared to microscopy and multiplex PCR. This study showed that the LAMP assay, targeting the actin gene, could be used to diagnose early infections of T. vaginalis. Thus, we have provided an alternative molecular diagnostic tool and a point-of-care test that may help to prevent trichomoniasis transmission and associated complications.
Trichomoniasis, which is caused by the protist, Trichomonas vaginalis, is one of the most common non-viral sexually transmitted infections, with an estimated prevalence of 276 million new infections worldwide each year [1,2]. T. vaginalis infection causes vaginitis and cervicitis, and is associated with serious complications, such as preterm birth, pelvic inflammatory disease, infertility, and an increased risk of co-infection with human immunodeficiency virus [3-9]. However, more than 50% of T. vaginalis infections in women are asymptomatic and remain undiagnosed [10]. Thus, early diagnosis and treatment of these infections, especially those that are asymptomatic, are imperative to alleviate this public health concern for women.
The most common clinical diagnostic test for T. vaginalis infection is microscopic examination of vaginal wet preparations (or “wet mount”); however, the sensitivity of wet mounts for detecting T. vaginalis is only 50-70% [11]. Recently, nucleic acid amplification tests, such as PCR and transcription-mediated amplification, using samples derived from vaginal swabs, have been developed. These methods show an improved sensitivity and specificity for the detection of T. vaginalis, compared to microscopic examination and culture, with sensitivity ranging from 64% to 100% [12,13].
Loop-mediated isothermal amplification (LAMP), which is a relatively straightforward and sensitive technique based on rapid DNA amplification under isothermal conditions, was recently developed to remove the need for sophisticated and expensive thermal cyclers [14]. LAMP involves the specific amplification of target DNA by a Bacillus stearothermophilus (Bst) DNA polymerase, which has a high strand displacement DNA synthesis activity. The assay is performed with a set of 4-6 primers that recognize independent regions of the target gene; this improves the specificity and the speed of the amplification. Herein loop-structured amplicons are formed, which results in a typical ladder-pattern of multiple bands imaged after DNA electrophoresis [15]. A positive reaction is easily determined by eye as turbidity [16] or fluorescence through the inclusion of fluorescent detection dyes, such as SYBR green or hydroxynaphthol, which can be detected using UV light [17,18]. These features allow the use of LAMP assays to detect many pathogenic organisms such as viruses, bacteria, fungi, and parasites [19-22].
The present study was undertaken to develop a LAMP assay detecting T. vaginalis and to validate the assay using vaginal swab samples obtained from females visiting the obstetrics and gynecology clinic with suspected T. vaginalis infections.
First, to select a target gene for the Trichomonas LAMP assay with high sensitivity and specificity, LAMP primers were designed specific to several candidate genes, such as Tvk [23-25], β-tubulin [26,27], and actin [28,29]; all of which were previously reported as PCR markers for the detection of T. vaginalis. Sequences were retrieved from GenBank (Tvk, GenBank no. XM_001579000; β-tubulin, GenBank no. L05468; actin gene, GenBank no. AF237734), and LAMP primers were designed based on previously described methods using the Primer Explorer program [30] (http://primerexplorer.jp/elamp4.0.0/index.html). LAMP primer sets for T. vaginalis consisted of F3 (forward outer primer), B3 (backward outer primer), FIP (forward inner primer), BIP (backward inner primer), LF (loop forward primer), and LB (loop backward primer) (Table 1). LAMP was performed in a Loopamp real-time turbidimeter (Realoop-30; Eiken Chemical Co., Ltd., Tokyo, Japan) for 90 min at 64˚C in a 25 μl mixture containing 40 pmol each of FIP and BIP, 5 pmol each of F3 and B3, 20 pmol each of LF and LB, 1.4 mM deoxynucleoside triphosphates, 0.8 M betaine, 1 μl of Bst DNA polymerase (New England Biolabs, Ipswich, Massachusetts, USA) with 2.5 μl of buffer (20 mM Tris–HCl, pH 8.8; 10 mM KCl; 10 mM (NH4)2SO4; 8 mM MgSO4; and 0.1% Tween 20), and 1 μl of template DNA. In our experiments, the actin gene was the only target amplified among the 3 candidate genes (data not shown) and was selected for further experiments. An optimal temperature and time for the LAMP reaction was determined using a cloned actin gene fragment (100 pg per reaction) under isothermal conditions at temperatures ranging from 60˚C to 65˚C for 120 min, by monitoring turbidity. Although amplification targeting of the actin gene was detected at all tested temperatures, a threshold value of absorbance (0.1) was reached fastest at 64°C (data not shown). Nonspecific amplification was not detected in the negative control (plasmid containing no insert) after at least 120 min of incubation. Thus, the subsequent LAMP reactions were conducted at 64˚C for 90 min.
To assess sensitivity, actin LAMP was conducted with serially diluted T. vaginalis genomic DNA ranging from 100 pg to 10 fg per reaction. This actin LAMP amplified the targeted region at each dilution from the highest concentration (1 ng per reaction) (Fig. 1A-C) to as little as 1 pg of T. vaginalis genomic DNA per reaction; all concentrations tested reached the threshold value of absorbance (0.1) within 40 min (Fig. 1A). As shown in Fig. 1B, a typical ladder pattern of amplified LAMP products was observed by agarose gel (2.0%) electrophoresis in lanes containing reactions performed with 1 pg to 1 ng of template DNA (Fig. 1B, lanes 1-4). To determine the specificity of the actin LAMP assay, the amplified LAMP products from the T. vaginalis genomic DNA were digested with HindIII. The sequence of the target region of actin contained this restriction site, and the expected fragments were generated (Fig. 1B, lane 5). Amplified products of the positive reactions were also visually detectable using the Loopamp fluorescent detection reagent (FD; Eiken Chemical Co., Tokyo, Japan) under UV light (Fig. 1C). Alternatively, DNA was extracted from approximately 1×102 parasites, serially diluted 10-fold to the equivalent of 100 to 0.01 parasites per reaction, and used as a template for the LAMP assays and PCR. The LAMP procedure amplified the DNA of parasites diluted to a concentration equivalent to 1 parasite per reaction (Fig. 1D). Similarly, PCR amplified the DNA of parasites at the equivalent dilution (Fig. 1E). Thus, the detection limit of the LAMP assay was consistent with that of PCR. The specificity of the T. vaginalis LAMP assay was evaluated with the genomic DNA of 3 known sexually transmitted disease-associated species (Candida albicans, Chlamydia trachomatis, and Neisseria gonorrhoeae) and 3 pathogenic but trichomoniasis-unrelated protozoan parasites (Cryptosporidium parvum, Entamoeba histolytica, and Giardia lamblia) and 1 unrelated bacterial species (Escherichia coli). As shown in Fig. 1F, color change (green) of the amplified LAMP products, with the Loopamp fluorescent detection reagent, was only observed for T. vaginalis genomic DNA. The T. vaginalis LAMP assay showed no detectable amplification of the other DNAs (brown).
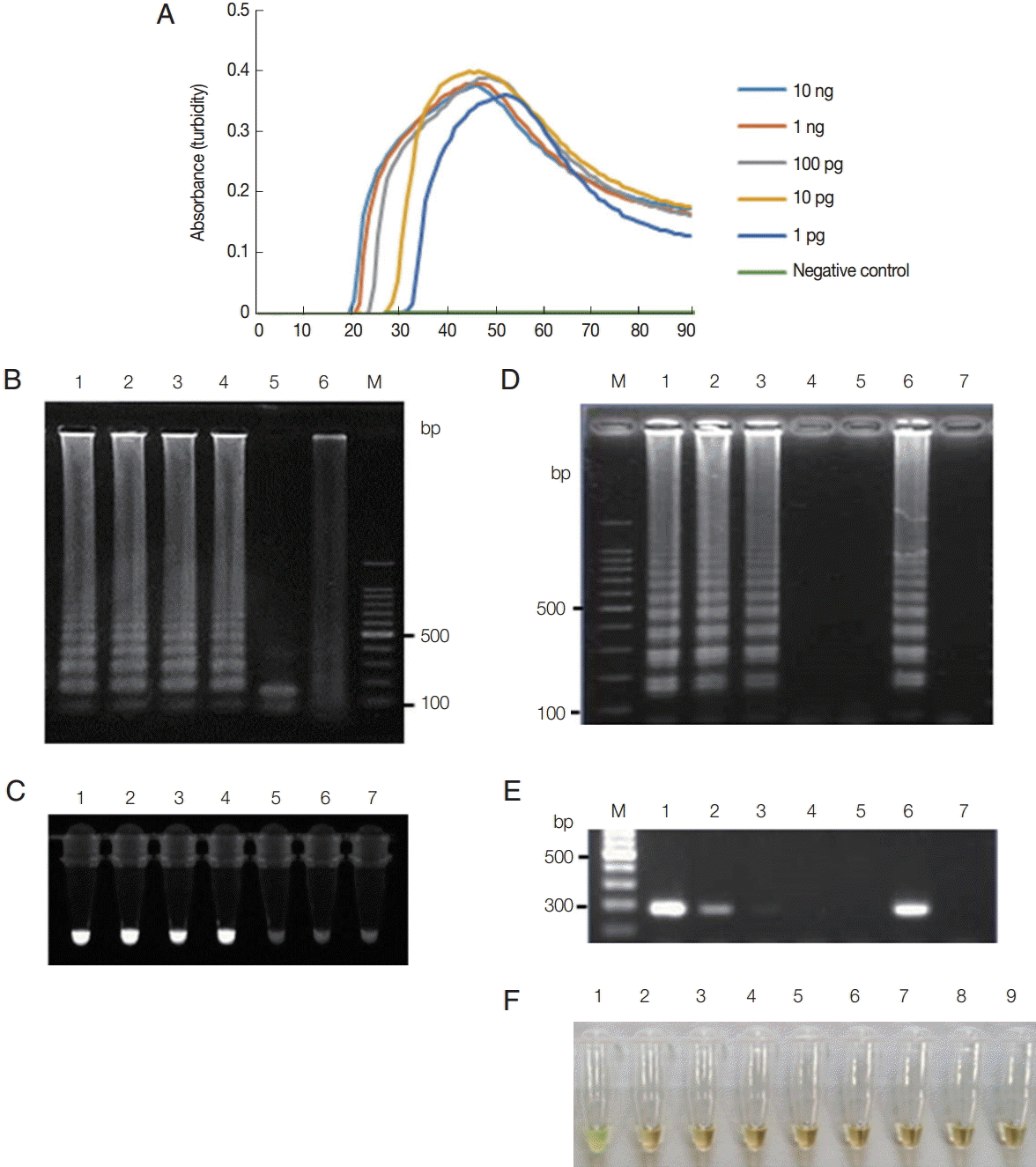
Functionality of T. vaginalis actin LAMP assays. (A) LAMP on 10-fold serial dilutions of T. vaginalis genomic DNA (10 ng to 1 pg per reaction) monitored by measuring absorbance. Distilled water was used as a negative control. (B) LAMP products were visualized by gel electrophoresis. Lane 1, 1 ng; lane 2, 100 pg; lane 3, 10 pg; lane 4, 1 pg of T. vaginalis genomic DNA; lane 5, LAMP product after HindIII digestion; lane 6, distilled water; lane M, 100-bp DNA marker. (C) LAMP products were visualized under UV light using the Loopamp fluorescent detection reagent. Lane 1, 1 ng; lane 2, 100 pg; lane 3, 10 pg; lane 4, 1 pg; lane 5, 100 fg; lane 6, 10 fg of T. vaginalis genomic DNA; lane 7, distilled water. (D-E) T. vaginalis at a density of 1×102 parasites/μl was serially diluted and tested using the LAMP assay (D) and PCR (E) with F3 and B3 primers (Table 1). Lane M, 100-bp DNA marker; lane 1, 100; lane 2, 10; lane 3, 1; lane 4, 0.1; lane 5, 0.01 parasite(s) per reaction; lane 6, positive control, 100 pg of plasmid DNA containing the LAMP targeting regions of actin gene; lane 7, distilled water. (F) Specificity of LAMP primers for detection of T. vaginalis assessed using template DNA from other microbial species. Lane 1, T. vaginalis; lane 2, Candida albicans; lane 3, Chlamydia trachomatis; lane 4, Neisseria gonorrhoeae; lane 5, Cryptosporidium parvum; lane 6, Entamoeba histolytica; lane 7, Giardia lamblia; lane 8, Escherichia coli; lane 9, human genomic DNA. LAMP products were visualized by a color change that was also observable by the naked eye under normal visible light.
To validate the T. vaginalis LAMP, vaginal swab specimens were obtained from 50 female patients, with clinical suspicion of trichomoniasis, visiting obstetrics and gynecology clinics. These samples were subjected to the LAMP assay and 3 alternative available methods of T. vaginalis detection, including microscopic examinations, PCR, and multiplex PCR. The study was approved by the Institutional Review Board of Kyungpook National University Hospital (IRB no. KNUH 2013-04-051). Wet mount microscopy was performed at the clinic and classified as positive when motile trichomonads were observed. Genomic DNA was extracted from vaginal swab specimens using a QIAamp DNA mini kit (Qiagen, Hilden, Germany) and the diagnosis of T. vaginalis infection was made using the results of multiplex PCR and PCR that targeted Tvk [27] in triplicate, as previously described [31]. In this study, PCR was used as a gold standard and the sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were calculated for each test (Table 2). The percentage sensitivities and specificities with 95% confidence intervals, differences in the sensitivities and specificities, and the degree of agreement were determined a previously described [31]. Of the 50 female patients, 12 (24%), 23 (46%), 40 (80%), and 43 (86%) were determined to be T. vaginalis positive using microscopy, multiplex PCR, PCR, and LAMP, respectively. All 40 positive specimens identified by PCR were positive using the LAMP assay. When PCR was used as the gold standard, the LAMP assay had the highest diagnostic sensitivity (P<0.001) and better agreement (kappa value: 0.789) with PCR than multiplex PCR and microscopy (Table 2). These findings indicate that the LAMP assay had a higher sensitivity for detection of samples infected with T. vaginalis compared to that of multiplex PCR and microscopic examination of wet mounts.
In this study, the LAMP assay was developed to detect T. vaginalis and was compared to currently available tests, including microscopic examinations, multiplex PCR, and PCR, using 50 vaginal swab specimens. Being a diagnostic test for trichomoniasis, microscopic examinations of vaginal swab samples are still routinely performed in primary clinics of South Korea. As shown in Table 2, microscopic examinations had the lowest sensitivity (30.0%) among the 4 tests performed in this study. This result is noteworthy since misdiagnosis of T. vaginalis, resulting from use of a less accurate diagnostic method, would result in a persistent infectious state, and may subsequently lead to serious T. vaginalis-associated complications and possible transmission of the infection. Among the currently available tests, PCR is still the most sensitive and specific method for T. vaginalis detection. However, the need for expensive laboratory equipment, trained specialists, and time-consuming procedures limit the usefulness of PCR in the primary clinic. In contrast, a LAMP assay does not require a thermal cycler and can be performed with minimal laboratory facilities, such as a heat block or a water bath, and yields results that can be assessed by eye. The straightforward sample preparation for molecular diagnosis reduces the risk of cross-contamination and the turnaround time for diagnosis, therefore, we prepared the crude cell lysates from vaginal swab specimens by direct heat-treatment [32].
LAMP assays and PCR performed on heat-treated lysates yielded the same results as those obtained using DNA extracted using a DNeasy tissue kit (data not shown). A T. vaginalis LAMP assay, based on 2-kb repeated DNA species-specific sequence has recently been developed [33] and evaluated for its sensitivity and specificity using spiked genital swabs and urine specimens. However, the goal of the present study was to select more efficient target DNA, through comparison of DNA amplification of candidate genes, and validate the assay using clinically obtained specimens. Moreover, the results of the actin LAMP assay using heat-treated samples were consistent with those of the LAMP assay using samples obtained from a commercial kit. Thus, this T. vaginalis LAMP assay could eliminate the need for DNA extraction, without compromising as say sensitivity, and reduce the time required to diagnose. Furthermore, individuals with low parasite density can provide reservoirs for transmission, and thus an early and accurate diagnostic tool such as the one described herein could help to prevent transmission of T. vaginalis infections.
Acknowledgements
This work was supported by a Biomedical Research Institute grant, Kyungpook National University Hospital (2013).
Notes
We have no conflict of interest related to this work.

