Molecular Phylogenetics of Trichostrongylus Species (Nematoda: Trichostrongylidae) from Humans of Mazandaran Province, Iran
Article information
Abstract
The present study was performed to analyze molecularly the phylogenetic positions of human-infecting Trichostrongylus species in Mazandaran Province, Iran, which is an endemic area for trichostrongyliasis. DNA from 7 Trichostrongylus infected stool samples were extracted by using in-house (IH) method. PCR amplification of ITS2-rDNA region was performed, and products were sequenced. Phylogenetic analysis of the nucleotide sequence data was performed using MEGA 5.0 software. Six out of 7 isolates had high similarity with Trichostrongylus colubriformis, while the other one showed high homology with Trichostrongylus axei registered in GenBank reference sequences. Intra-specific variations within isolates of T. colubriformis and T. axei amounted to 0–1.8% and 0–0.6%, respectively. Trichostrongylus species obtained in the present study were in a cluster with the relevant reference sequences from previous studies. BLAST analysis indicated that there was 100% homology among all 6 ITS2 sequences of T. colubriformis in the present study and most previously registered sequences of T. colubriformis from human, sheep, and goat isolates from Iran and also human isolates from Laos, Thailand, and France. The ITS2 sequence of T. axei exhibited 99.4% homology with the human isolate of T. axei from Thailand, sheep isolates from New Zealand and Iran, and cattle isolate from USA.
INTRODUCTION
Trichostrongylus species are parasitic nematodes of the small intestine of ruminants, rodents, pigs, horses, birds, and humans with a worldwide distribution [1,2]. There are more than 30 species of Trichostrongylus most of which are parasites of herbivores. At least 10 species have been reported from humans where people and herbivorous animals are in close contact [3,4]. Human infections occur mainly via ingestion of filariform larvae from contaminated vegetables or water or rarely by penetrating through the skin [3]. It is common locally in many countries, including Iran, Iraq, Egypt, Ethiopia, Laos, Thailand, South Korea, China, Japan, and United States [5]. Most human infections cause no clinical symptoms and symptomatic individuals may present with abdominal pain, diarrhea, and eosinophilia [6,7].
Classification of Trichostrongylus spp. by conventional morphological methods is relatively reliable on Trichostrongylus males. However, these methods are laborious and cannot be relevant to recognize female worms [8]. Finding the characteristic eggs of Trichostrongylus in stool samples is a routine diagnostic method, but is not helpful to differentiate the species [8].
In recent years, PCR-based techniques are applied for species identification and phylogenetic analysis of Trichostrongylus nematodes in ruminants worldwide [9–13]. There are also a few studies on identification and genetic characterization of Trichostrongylus samples from humans, such as third-stage larvae [3,7,14] and eggs in stool samples [15], by the detection of DNA. Investigations have shown ribosomal DNA sequencing (particularly ITS2 region) as a useful tool for differentiation of Trichostrongylus species and analysis of genetic variations and phylogenetic relationships [3,9,10,13,15]. In Iran, using morphological methods, human infections with 8 species of Trichostrongylus were reported, including T. capricola [16], T. lerouxi [17], T. orientalis, T. vitrinus, T. axei, T. colubriformis, T. probolurus, and T. skrjabini [4]. Among those, T. orientalis and T. colubriformis were detected more frequently in regions where the prevalence of infection was high [4]. Above mentioned species were also reported in ruminants in Iran, by morphological [4,18,19] and molecular methods [9], indicating zoonotic potential of those species. However, phylogenetic studies on human isolates of trichostrongyliasis in Iran are lacking. Therefore, we tried the molecular phylogenetic analysis based on ITS2 region of ribosomal DNA with fecal egg samples of Trichostrongylus, collected from residents in Mazandaran Province, northern Iran.
MATERIALS AND METHODS
Study area and sample collection
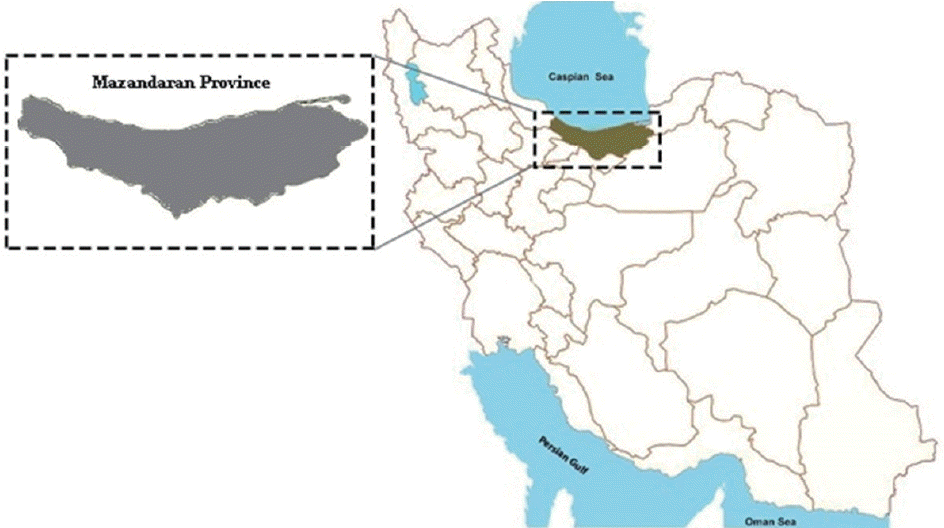
Mazandaran Province is situated on the southern coast of the Caspian Sea in the north of Iran (53°6′ E, 36°23′N) (Fig. 1). It has a humid weather with annual average rainfall of 977 mm. This province is geographically divided into 2 parts; the coastal plains and the mountainous regions [20]. Most rural residents are farmers, and domestic animal husbandry can expose them to zoonotic parasites, such as Trichostrongylus. This province has previously been known prevalent for Trichostrongylus species, both in domestic animals [4] and humans [4,15]. In this study, 7 human trichostrongyliasis cases were detected by formalin-ether concentration technique during a study on evaluation of molecular and parasitological methods for the diagnosis of strongyloidiasis in fecal samples [21]. They were 5 males and 2 females with ages ranging between 23 and 57 years old, residing in Mazandaran Province, northern Iran. For extraction of genomic DNA, fecal samples of the patients were kept in 70% ethanol at room temperature.
Molecular and phylogenetic analysis
DNA of the samples was extracted by using in-house (IH) method as described by Repetto et al. [22] and modified by Sharifdini et al. [21]. Briefly, 1 g of stool samples was diluted in 10 ml of PBS and was subjected to 5 cycles of freezing and thawing. Next, 500 μl of the PBS diluted stools were incubated overnight with 500 μl GTES buffer at 37°C, followed by 3 times freezing–thawing. Then, 200 mg of glass beads were added and shaken rigorously for 5 min. The suspension was incubated for 12 hr in nematode lysis buffer at 37°C. Then, the samples were extracted with phenol-chloroform-isoamyl alcohol (25:24:1), and DNA was precipitated with an equal volume of isopropanol and 1 ml of 100% ethanol, respectively. The pellet was washed with 300 μl of 70% ethanol, dried and eluted in 100 μl of TE buffer and stored at −20°C until PCR amplification.
The ribosomal DNA internal transcribed spacer 2 (ITS2) region was amplified by forward (NC1: 5-ACGTCTGGTTCAGGGTTGTT-3) and reverse (NC2: 5-TTAGTTTCTTTTCCTCCGCT-3) primers [23]. The PCR reactions were performed in a final reaction volume of 30 μl containing 15 μl of PCR mix which included 1.25 U Taq DNA polymerase, 200 μM of dNTPs, and 1.5 mM MgCl2 (2x Master Mix RED Ampliqon, Copenhagen, Denmark), 10 pmol of each primer, and 4 μl of DNA sample. The PCR program was an initial denaturation step at 95°C for 6 min followed by 35 cycles of 94°C for 45 sec (denaturation), 60°C for 90 sec (annealing), and 72°C for 60 sec (extension), followed by a final extension at 72°C for 5 min.
The PCR products were run on a 1.5% agarose gel. DNA sequencing was performed using an ABI 3130xl platform (Applied Biosystems, Foster City, California, USA). The sequence results were edited and analyzed by the Geneious software (www.geneious.com) and compared with sequences deposited in GenBank by BLAST program (http://www.ncbi.nlm.nih.gov/). A phylogenetic tree was constructed using the maximum likelihood method based on the Tamura 3-parameter model, and pairwise comparisons were determined of the level of sequence differences within and among species using MEGA 5.0 software. Bootstrap analysis was done based on 1,000 replications.
RESULTS
Microscopical examination of all 7 stool sedimentations revealed the presence of Trichostrongylus eggs with their morphological characteristics, including elongated oval shape, and slight tapering at one end. However, alignments of ITS2 sequences of these isolates separated them as T. colubriformis (Fig. 2A) and T. axei (Fig. 2B).
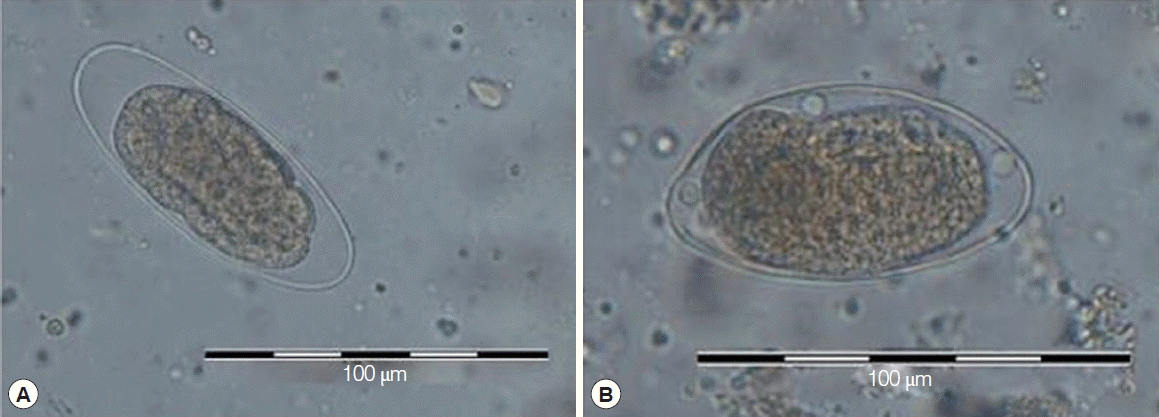
Light microscope view of Trichostrongylus colubriformis (A) and Trichostrongylus axei (B) eggs in stool samples of infected people showing morphological characteristics, including elongated oval shape and slight tapering at one end.
All samples were successfully presented amplification of about 328 bp for the ITS2 gene. Comparisons of the sequences from these isolates with other available reference sequences in GenBank, using BLAST system, revealed that 6 isolates had high similarity (more than 95%) with T. colubriformis, and the other one had high homology with T. axei. These sequences were deposited in GenBank database (accession nos.: KP663663, KP663664, KF989494, KF989495, KF989496, and KF989497 for T. colubriformis and KF840722 for T. axei).
The multiple alignments of ITS2 sequences of Trichostrongylus spp. obtained in this study are available in GenBank. Intra-specific variation within isolates of T. colubriformis and T. axei amounted to 0–1.8% and 0–0.6%, respectively; while inter-specific sequence differences among Trichostrongylus nematodes were significantly higher, being 1.8–7.0%. Based on these variations, 21 isolates of T. colubriformis and 5 isolates of T. axei were classified into 5 and 2 haplotypes for ITS-2, respectively (Fig. 3). Trichostrongylus species of the present study were in a cluster with the relevant reference sequences from previous studies (Fig. 4).
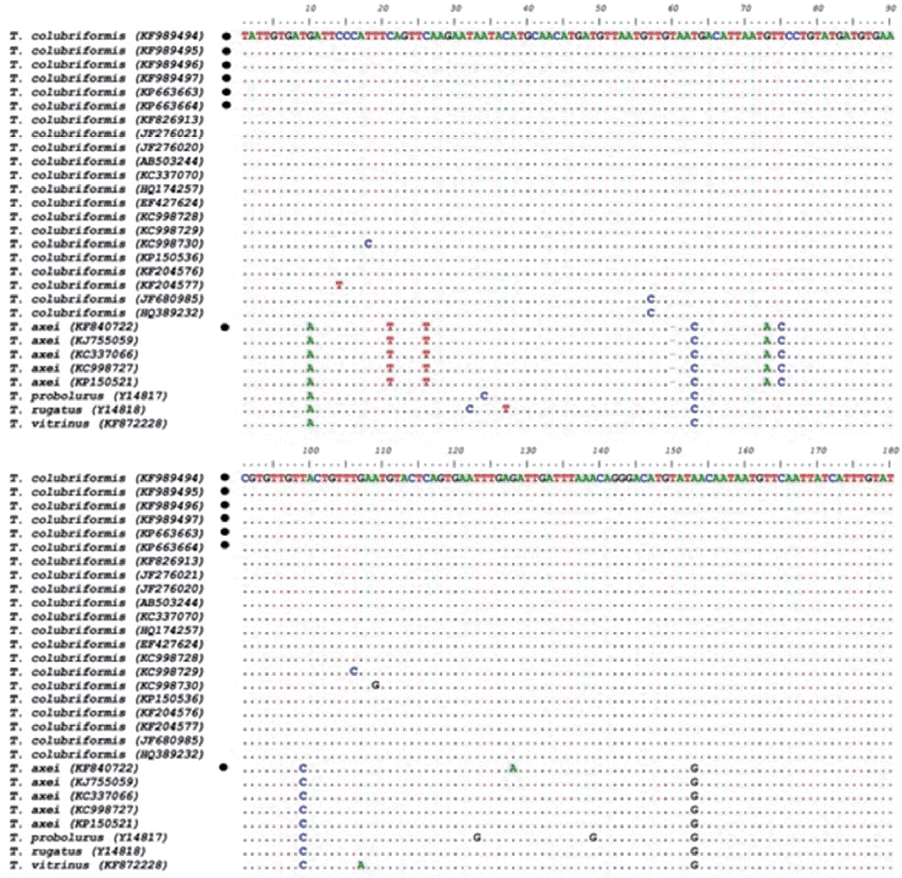
Alignments of the ITS2 sequences for all Trichostrongylus isolates in the present study and reference sequences retrieved from GenBank. Sequences with circle mark were inferred from this study.

Phylogenetic tree of isolates of Trichostrongylus spp. obtained in this study (▲) and reference sequences retrieved from GenBank based on ITS2 nucleotide sequences and constructed using the Tamura 3-parameter model in MEGA software version 5. GenBank sequences of Trichostrongylus spp. included with Necator americanus as an out group. Iran (IRN), France (FRA), Thailand (THA), Laos (LAO), Russia (RUS), Malaysia (MYS), United States (USA), Ireland (IRL), Australia (AUS), and New Zealand (NZL) are represented with country codes (ISO 3166-1 a-3 codes).
DISCUSSION
Infection with species of Trichostrongylus is common among herbivores in most parts of Iran. T. colubriformis [4,18,19,24,25], T. vitrinus [4,18,19,24,25], T. axei [4], T. capricola [4,19], T. probolurus [4,18,19,24,25], T. longispicularis [19], T. orientalis [4], T. lerouxi [26], T. skrjabini [4], and T. hamatus [25] were reported in different animals, such as sheep [4,18,19], goat [4,19], cattle [4,19], camel [24,25], and buffalo [4,19] using morphology. Predominant species among different herbivores in most parts of the country like Mazandaran Province are T. colubriformis, T. vitrinus, and T. axei [4]. Considering human infections, high prevalence of infection [4], as well as variety of Trichostrongylus species including T. orientalis [4], T. colubriformis [4], T. vitrinus [4], T. axei [4], T. capricola [16], T. probolurus [4], T. skrjabini [4], and T. lerouxi [17], has been reported from Iran, back to 1970’s, indicating T. colubriformis and T. orientalis as predominant species [4]. Recently, among human geohelminths, the prevalence of some species especially Ascaris lumbricoides and hookworms are sharply declined [27]; however, Trichostrongylus spp. [15,28,29] and Strongyloides stercoralis [21,29,30] are more frequently reported due to zoonosis of the former and ability of auto infection of the latter parasite, respectively.
In this study, in spite of the availability of few human Trichostrongylus samples (n=7), using ITS2 sequence analysis, 6 of them were determined as T. colubriformis, and the other one as T. axei. This result is compatible with the result of a recent molecular study on human trichostrongyliasis in Mazandaran Province, in which T. colubriformis was accounted as the most probable common species of Trichostrongylus in humans [15]. Human trichostrongyliasis is caused by using animal feces as a fertilizer for agriculture and gardening [31]. In the north of Iran, many domestic animals, such as sheep, goats, and cows graze almost freely around, and contamination risk of vegetables in the fields with animal feces is high. Additionally, in gardening practices, use of fresh sheep and cattle residues as fertilizer is common. Therefore, T. colubriformis is among more frequent geohelminths of humans in the study area due to its high zoonotic capability and its high prevalence in domestic animals [4]. While, the prevalence of T. orientalis, similar to human hookworms and also A. lumbricoides, has decreased due to not using human nigh soil as fertilizer in the study area, as in most other parts of the country.
Phylogenetic sequence analysis is a useful tool to gain information on an organism’s evolutionary relationships. The existence of genetic variation among Trichostrongylus nematodes has been confirmed previously [9,10,32]. However, only a few studies have analyzed molecular-phylogenetic characterization of human trichostrongyliasis [3,14,33]. This study is the first phylogenetic analysis of Trichostrongylus species from humans in Iran. BLAST analysis of the isolates indicated that sequences of all 6 T. colubriformis had 100% homology with each other and with the previously registered sequences from the human (KF826913), sheep (JF276021), and goat (JF276020) in Iran. Among GenBank sequences of T. colubriformis, 1 sequence from sheep in Iran (HQ389232) had 1 nucleotide difference in ITS2 with that of T. colubriformis in this study. The latter sequence, in phylogenetic tree, was placed in 1 group along with a sheep isolate of T. colubriformis sequence from Ireland (JF680985). Sequences of T. colubriformis isolates in present study also presented 100% homology with T. colubriformis in humans from France (HQ174257), Thailand (KC337070), and Laos (AB503244). These sequences also showed 100% similarity with T. colubriformis from sheep (EF427624 and KC998728), goat (KF204576), and cattle (KP150536) in other countries. For these sequences, 99.4% similarities were obtained with T. colubriformis isolates from sheep in Ireland (JF680985), sheep in New Zealand (KC998729) and goat in Malaysia (KF204577). T. colubriformis sequences of this study presented 98.8% homology with the sequences of T. colubriformis from sheep in New Zealand (KC998730).
Human infections with T. axei were reported among the literatures in Iran [4] and Italy [34]. Recently, T. axei was also found in humans in Thailand [3] and Iran [15] by molecular techniques. The ITS2 sequence of T. axei in this study exhibited 99.4% homology with the human isolate of T. axei from Thailand (KC337066), sheep isolate from New Zealand (KC998727), sheep isolate from Iran (KJ755059), and cattle isolate from USA (KP150521). T. axei of the current study (KF840722) has 1 nucleotide difference with the reference sequences of T. axei from Iran and other countries.
To sum up, T. colubriformis was found to be the most probable dominant human species in the study area, but further investigation with higher sample size is recommended. This species is also the most possible cause of human trichostrongyliasis infection in Laos [14], Thailand [3], and France [7]. It might also be the main zoonotic species of Trichostrongylus in some other parts of the world which have not been investigated yet.
Current phylogenetic analysis clarified the relation of human Trichostrongylus species from an endemic area of trichostrongyliasis in Iran and those of human and animal species of Trichostrongylus registered in GenBank. Based on pairwise comparisons, there was 100% homology among all 6 ITS2 sequences of T. colubriformis in the present study and most previously registered sequences of T. colubriformis from human and herbivores animals. High zoonotic capacity of this species is probably one of the main reasons of current higher occurrence of human trichostrongyliasis in the study area than that of human hookworms which are not zoonotic. Comparative characterization of Trichostrongylus species based on molecular approach from human and different livestock in every endemic area, using several gene targets, will be interesting and can be beneficial to understand the rate of zoonosis of each species.
ACKNOWLEDGMENTS
The financial support of this study was partly provided by the Deputy of Research, Tehran University of Medical Sciences, Tehran, Iran, through grant no. 91-01-160-17294. The authors would like to thank all people who have contributed to this research, especially Mrs. B. Kamranrashani from the School of Public Health, Tehran University of Medical Sciences and T. Hesari for their kind help.
Notes
CONFLICT OF INTEREST
The authors declare that they have no conflicts of interest.