Infection Status of Hospitalized Diarrheal Patients with Gastrointestinal Protozoa, Bacteria, and Viruses in the Republic of Korea
Article information
Abstract
To understand protozoan, viral, and bacterial infections in diarrheal patients, we analyzed positivity and mixed-infection status with 3 protozoans, 4 viruses, and 10 bacteria in hospitalized diarrheal patients during 2004-2006 in the Republic of Korea. A total of 76,652 stool samples were collected from 96 hospitals across the nation. The positivity for protozoa, viruses, and bacteria was 129, 1,759, and 1,797 per 10,000 persons, respectively. Especially, Cryptosporidium parvum was highly mixed-infected with rotavirus among pediatric diarrheal patients (29.5 per 100 C. parvum positive cases), and Entamoeba histolytica was mixed-infected with Clostridium perfringens (10.3 per 100 E. histolytica positive cases) in protozoan-diarrheal patients. Those infected with rotavirus and C. perfringens constituted relatively high proportions among mixed infection cases from January to April. The positivity for rotavirus among viral infection for those aged ≤ 5 years was significantly higher, while C. perfringens among bacterial infection was higher for ≥ 50 years. The information for association of viral and bacterial infections with enteropathogenic protozoa in diarrheal patients may contribute to improvement of care for diarrhea as well as development of control strategies for diarrheal diseases in Korea.
INTRODUCTION
Acute diarrhea is one of the most common diseases worldwide, and causes approximately 2.5 million children death annually [1-3]. In the Republic of Korea (= Korea), the mortality rate in children younger than 5 years was 0.3% [4]. The protozoan parasite, including Cryptosporidium parvum, Giardia lamblia, and Entamoeba histolytica, have been under constant surveillance in developed countries, including USA, UK, and Japan due to their potential for causing water- and food-borne diarrheal outbreaks [5-7]. Also water- and food-borne viruses have been implicated with acute gastrointestinal diseases in humans, and these enteric viruses have occasionally been identified as the etiologic agents of waterborne disease outbreaks [8-11]. Rotavirus has been shown to be responsible for 25% to 46% of diarrhea episodes among hospitalized children < 5 years old in Korea, 2002-2004 [12].
The Korea National Institute of Health (KNIH) has surveyed pathogenic agents for hospitalized diarrheal patients every year from 2003 across the nation. This surveillance program examined 10 species of enteropathogenic bacteria, including Salmonella spp., Shigella spp., enterotoxigenic Escherichia coli, Vibrio parahaemolyticus, Yersinia enterocolitica, Staphylococcus aureus, Clostridium perfringens, Campylobacter jejuni, Listeria monocytogenes, and Bacillus cereus, and 4 species of viruses, including rotavirus, adenovirus, astrovirus, and norovirus in the stools of around 30,000 patients each year. The examination of major water- and food-borne protozoa (C. parvum, G. lamblia, and E. histolytica) causing gastrointestinal diseases was added to the program in 2004.
Global studies have identified and analyzed the pathogenic bacteria and viruses and protozoa that cause diarrhea, but there have been few studies in Korea. Furthermore, there is especially no study of mixed infections of viruses and bacteria in relation to protozoan infections in diarrheal patients. In this study, we evaluated the infections of acute gastrointestinal protozoa, bacteria, and viruses in hospitalized patients who had a major symptom of diarrhea. This study is the first to evaluate the infections of parasitic protozoa and their mixed infections with acute gastrointestinal pathogenic bacteria and viruses in patients who were hospitalized with a major symptom of diarrhea across the nation of Korea.
MATERIALS AND METHODS
Surveyed areas and stool samples
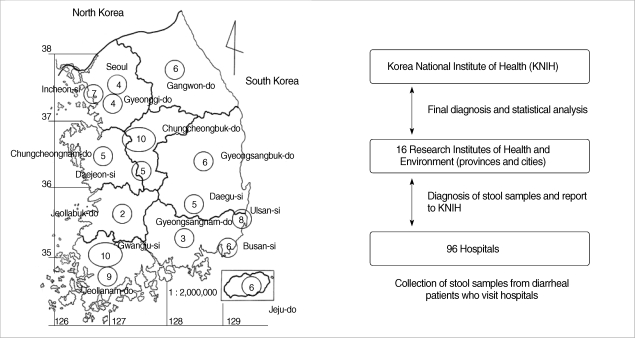
The survey was carried out as a part of the national program for control of diarrheal diseases from 2004 to 2006. The prevalence study was planned and implemented by the Department of Malaria and Parasitic Diseases, KNIH (Korea National Institute of Health). RIHE (Research Institute of Health and Environment) in 16 cities and provinces collected stools from 76,652 diarrheal patients who were hospitalized in 96 hospitals (Fig. 1). The diarrheal patients in this study were defined as those having diarrhea 3 or more times per day in watery or loose form accompanied by vomiting and abdominal pain. One gram or 1 loop of each stool sample was tested.
Protozoa
Cysts or oocysts of C. parvum, G. lamblia, and E. histolytica in the stool samples were identified with an enzyme immunoassay kit (Ridascreen, Darmstadt, Germany). Briefly, approximately 1 g of stool sample was suspended in 3.5 ml of distilled water and then filtered through moistened gauze. The filtrated liquid was centrifuged at 1,660 g for 3 min at 4℃. The supernatant was discarded and the pellet was tested according to the manufacturer's protocol. The absorbance was measured at 450 nm. Negative and positive controls were determined at absorbance values lower than 0.2 and higher then 0.8, respectively.
Bacteria
To identify bacteria (Salmonella spp., Shigella spp., enterotoxigenic E. coli (EPEC, ETEC, EAEC, STEC, EIEC), V. parahaemolyticus, Y. enterocolitica), a loop of each stool sample was directly inoculated into 3 ml of Luria Burtani (LB; Sparks, Maryland, USA) broth and incubated overnight at 37℃. The enriched broth culture was centrifuged at 23,000 g for 1 min. The pellet was heated to 100℃ for 10 min and spun down. A total of 5 µl of the supernatant was used in PCR. To detect pathogen-specific target gene, PCR was performed using the primers listed in Table 1. In brief, PCR was carried out in 50 µl with 2 U of DNA Taq polymerase (Takara, Tokyo, Japan) in a thermal cycler (PTC-100, MJ Research, Watertown, Massachusetts, USA). To further isolate, 8 different selective agar plates were used; MacConkey agar (BD) for E. coli, Salmonella, and Shigella spp, thiosulfatecitrate-bile salts-sucrose (TCBS, Oxoid, Basingstoke, Hampshire, UK) agar for Vibrio spp., mannitol-salt agar (MSA, BD) for S. aureus (toxins A, B, C, D, and E), tryptose-sulfite-cycloserine (TSC, Oxoid) for C. perfringens (alpha toxin, enterotoxin, or nontoxin), blood-free selective agar base (CCDA, BD) for C. jejuni, Listeria selective agar (LSA, Oxoid) for L. monocytogenes, cefsulodin-irgasan-novobiocin (CIN, Oxoid) for Y. enterocolitica, and mannitol-egg yolk-polymixin (MYP, Oxoid) for B. cereus (nemolysin BL-enterotoxin, nonhemolytic enterotoxin). One well-isolated colony from each bacteria culture was inoculated into 5 ml of 0.85% NaCl medium (pH 5.5 to 7.0). A humid atmosphere was used, and the API kit (Biomerieux, Marcy l'Etoile, France) was used as directed by the manufacturer to identify the organisms. The detection of Salmonella spp., pathogenic E. coli, C. perfringens, S. aureus, and B. cereus was considered positive regardless of the toxin in results.
Virus
Rotavirus, adenovirus, and astrovirus were analyzed using a VIro-Capture kit (Bioincell, Houston, Texas, USA), VIro-Capture kit (Bioincell), and IDEIA kit (DAKO, Glostrup, Denmark), respectively, according to the manufacturer's protocol. Norovirus was identified by reverse-transcription PCR [11].
Statistical analysis
Categorical variables were compared using the chi-square test. Logistic regression analysis was performed to examine factors that were associated with protozoa, viruses, and bacteria. The multi-variable models were fit adjusting for the variables (gender, age group, and area). Statistical significance (P < 0.05) was defined at the 95% confidence interval. SAS software (version 9.1) was used for all statistical analyses.
RESULTS
Positivity for protozoa, viruses, and bacteria
We examined 76,652 stool samples from diarrheal patients whose chief complaint was diarrhea. Positivity of our investigated 3 protozoa, 4 viruses, and 10 bacteria was 28,237 diarrheal patients (3,684 per 10,000 individuals), and their positivity was found to be 129 (95% confidence interval [CI], 121-137), 1,759 (95% CI, 1,732-1,785), and 1,797 (95% CI, 1,769-1,824) per 10,000 persons, respectively; it was lower for protozoa than for viruses and bacteria (P < 0.0001). Males showed a higher prevalence rate than females (55%, 33,166 persons) (P < 0.0001), but that was not different according to gender for parasites (P = 0.2334), viruses (P = 0.2206), and bacteria (P = 0.1198). We divided the age groups of diarrheal patients into ≤ 5 year, 6-9 year, 10-49 year, and ≥ 50 year groups. The highest number of diarrheal patients was ≤ 5 year group, 13,674 of 76,652 (4,240 per 10,000 individuals). The positivity for protozoa was significantly high in ≤ 5 year group compared to 6-9 year group (P = 0.0115). The positivity for viruses for those aged ≤ 5 years (P < 0.0001) was significantly high, while that for bacteria was low (P < 0.0001). The area distribution did not affect the positive rate for protozoan infections (P = 0.0601), but the adjusted odds ratio (AOR) (1.5, P < 0.0001) for viruses and for bacteria (1.1, P < 0.0001) was relatively high in provincial areas (Table 2).
Age-related positivity
Table 3 lists the positivity for protozoa, viruses, and bacteria among diarrheal patients by age group. Protozoan and viral infections were evident in ≤ 5 year group. The number of protozoan infections was 854 of 24,240 individuals (350 per 10,000 individuals), and 25.1% of C. parvum, 55.0% of G. lamblia, and 20.0% of E. histolytica positive rates were observed. The positivity for C. parvum was significantly higher at ≤ 5 years than at 10-49 years (P < 0.0001) and ≥ 50 year (P = 0.0033). The positivity for G. lamblia was higher than that for C. parvum and E. histolytica across all age groups (P < 0.0001). The positivity for norovirus, rotavirus, and adenovirus was low in old individuals, while that for astrovirus was high in those aged ≥ 50 year than in younger age groups (6-9 years and 10-49 years). In the case of bacteria, the positivity for Salmonella spp. was significantly high at 6-9 years compared to other age groups (P < 0.0001). The positivity for Shigella spp. was high in old age groups, but there was no significant difference (P = 0.7108). The positivity for V. parahaemolyticus, C. jejuni, and E. coli was highest at 10-49 years, and that of C. perfringens was highest at ≥ 50 years. The positivity for B. cereus was high across all ages (P = 0.0057), while that of S. aureus was significantly high at ≤ 5 years compared to other age groups (P < 0.0001). No significant difference was observed for L. monocytogenes and Y. enterocolitica due to their low positivity in each age group. The positivity for S. aureus was the highest in those aged ≤ 5 years, while that of C. perfringens was highest in other age groups.
Mixed-infections
The probability of mixed infection with viruse or bacteria among protozoan-infected cases (n = 987) was investigated (Table 4). A total of 535 patients of protozoan diarrhea was mixed-infected with viruses (311, 58.1%) or bacteria (224, 41.9%), respectively. Mixed infections with viruses were prominent in C. parvum diarrheal patients, and opposite results were observed in G. lamblia and E. histolytica infections. These 3 kinds of protozoa were mainly mixed-infected with rotavirus and C. perfringens. The positivity (probability) of mixed infection with C. parvum and rotavirus was the highest, 29.5 (95% CI, 23.9-35.1) per 100 C. parvum-infected persons, while that of mixed infection with E. histolytica and C. perfringens was 10.3 (95% CI, 6.0-14.6) per 100 E. histolytica-infected cases.
Seasonal prevalence
In Korea, cold winter months are from November to March, dry and mild spring months are from April to May, hot and moist summer months with sudden showers are from June to August, and dry and cool autumn months are September and October. The seasonal trends of mixed infections with protozoa and bacteria or viruses were analyzed (Fig. 2). The proportion of mixed infections was relatively lower in G. lamblia infection than in C. parvum or E. histolytica infection. The seasonal trend from January to April was higher with rotavirus and C. perfringens than the other pathogens.
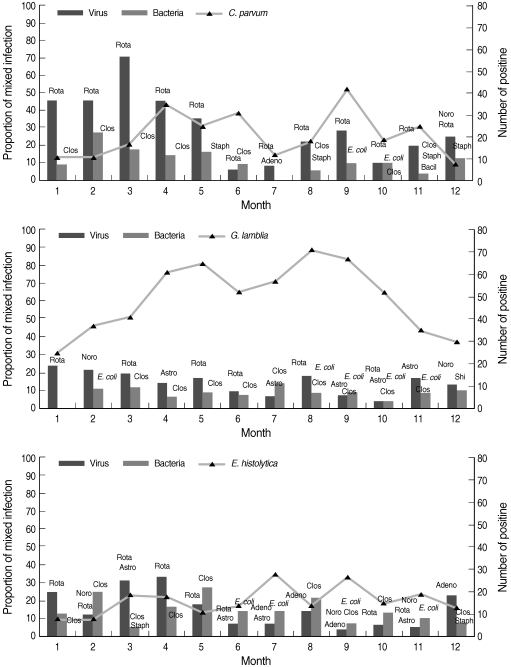
Seasonal patterns of mixed infections with protozoa. Noro, norovirus; Rota, rotavirus; Adeno, enteric adenovirus; Astro, astrovirus; Clos, Clostridium perfringens (alpha toxin, enterotoxin, or nontoxin); Staph, Staphylococcus aureus (toxins A, B, C, D, and E); E. coli, pathogenic E. coli (STEC, EAEC, EPEC, ETEC, and EIEC); Bacil, Bacillus cereus (hemolysin BL-enterotoxin, nonhemolytic enterotoxin); Shi, Shigella flexneri.
DISCUSSION
This is the first systematic study in which the distributions of various enteropathogens and mixed infections related to diarrhea are described in Korea. Our findings highlight the importance of diarrheal diseases associated with protozoan infections. The parasitic protozoa might infect the intestinal mucosa and elicit transient or chronic diarrhea, which should be considered as one of the most important opportunistic gastrointestinal pathogens. C. parvum and G. lamblia might constitute the principal candidates for endemic transmission due mainly to their ubiquitous presence in drinking water and food contents, and partly to their high resistance to relevant environmental factors and chemical disinfecting procedures [13,14]. C. parvum was highly mixed-infected with rotavirus that often infects pediatric hospitals, which has a high infection rate amongst those aged ≤ 5 years, whose immune systems may be weaker than normal [15,16]. In developing countries, rotavirus infections in the gastrointestinal tract constituted a major cause of childhood death, especially in children younger than 5 years old. It has been responsible for approximately half a million deaths per year [9,10,17,18]. In the present study, C. parvum infection should be considered among viral mixed infections, which is important to consider in therapies applied to hospitalized children patients.
In Korea, most diagnoses and treatments for diarrheal patients have ignored protozoan infections because most clinicians focused their diagnosis on the pathogenic viruses and bacteria. In this situation, it is possible that the unknown pathogens accounting for outbreaks of over 12% of food-borne diseases contain manifold protozoa [19]. Even though the infection rate of gastrointestinal protozoa is not so high (1.0%), gastrointestinal infections with protozoan parasites should not be overlooked.
C. perfringens, which is a spore-forming bacterial indicator of fecal contamination and an important food-borne diarrheal causative agent, was found to be highly mixed-infected with protozoa. The infection rate for C. perfringens was the highest in those aged ≥ 50 years. C. perfringens was found in 7.6% of 662 stool samples from diarrheal patients, whereas infection rates of Salmonella spp., S. aureus, and Y. enterocolitica were 5.4%, 3.0%, and 0.2%, respectively [20]. Most pediatricians prescribe antibiotics in less than 20% of patients with acute gastroenteritis infection, while most physicians routinely prescribe antibiotics in Korea [21].
One question arising from this study is whether mixed infection with protozoa is more likely to induce serious diarrhea. As usual, single infection caused more severe symptoms compared to mixed infections in a patient. It is also depending on the strength of the pathogenic strain as well as the infectious species involved [22-25]. In most cases, detailed information about clinical symptoms and the infectious density of diarrheal patients was obscured in this study. Rotavirus and C. perfringens were, however, highly mixed-infected with enteropathogenic protozoa in diarrheal patients. Identification of etiologic agents of acute gastrointestinal infections has important implications for their management and prevention [26]. A better understanding of the determinants of diarrheal medical care and drugs prescribed is clearly required to address the burden of acute gastrointestinal infection in Korea. The novel information obtained in the present study regarding infection and mixed-infection status of enteropathogenic bacteria and viruses with protozoa in diarrheal patients will help in further isolation and treatment of diarrheal diseases.
ACKNOWLEDGEMENTS
We express our gratitude to our colleagues in the RIHEs of the 16 cities and provinces for their collection of stools from diarrheal patients. They also performed laboratory examinations to detect 3 protozoan parasites, 10 bacteria, and 4 viruses. This work was supported by a grant from the National Institute of Health (NIH-091-4800-4851-300), National Research and Development Program, Ministry of Health and Welfare, the Republic of Korea.